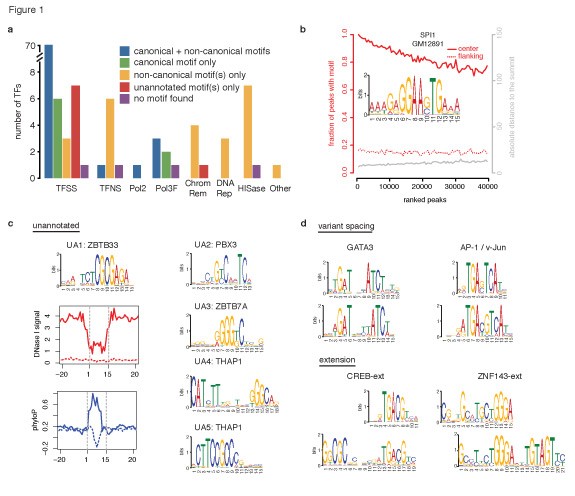
Figure 1 from Predicting transcription factor binding motifs from DNA- binding domains, chromatin accessibility and gene expression data | Semantic Scholar
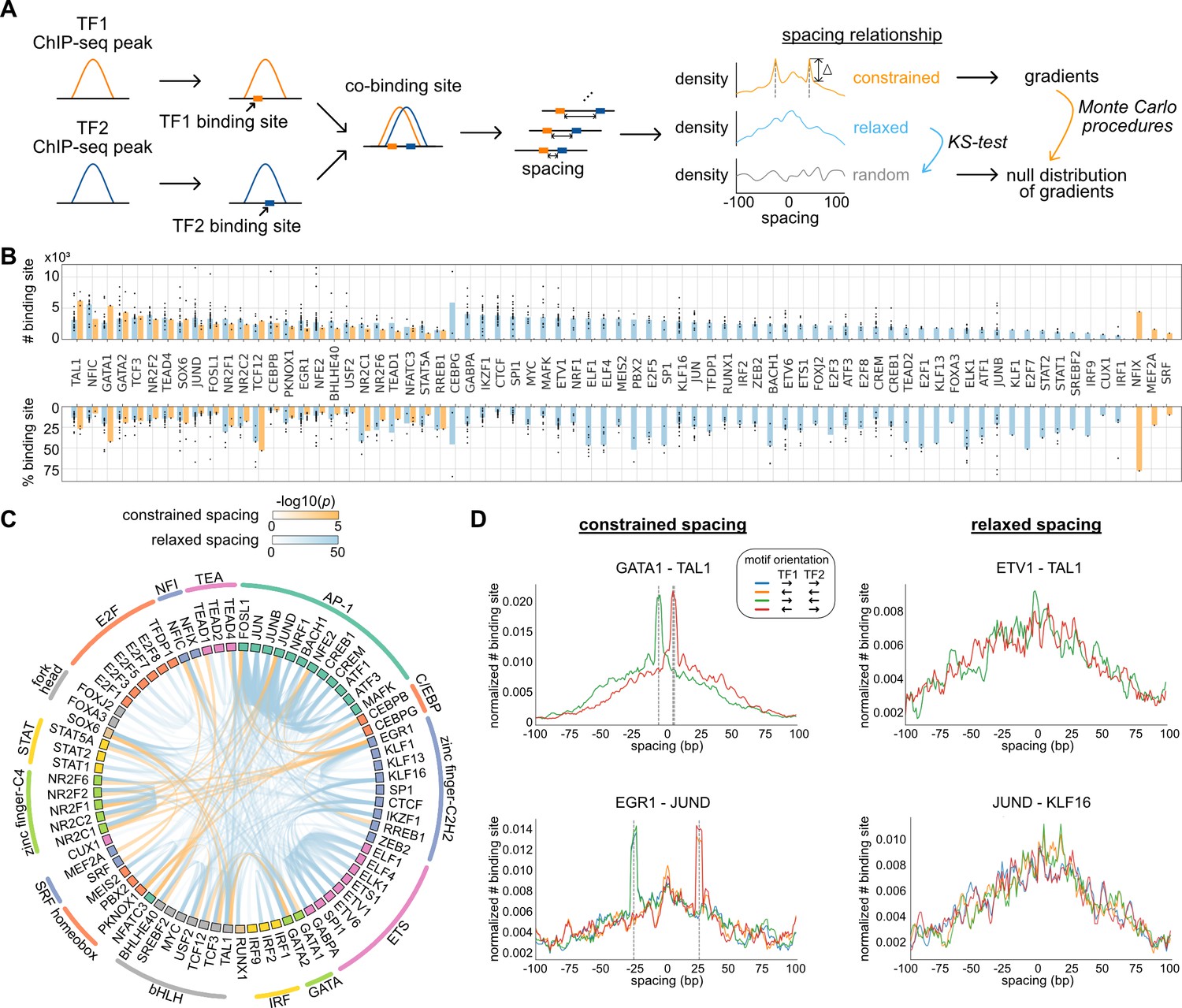
Systematic analysis of naturally occurring insertions and deletions that alter transcription factor spacing identifies tolerant and sensitive transcription factor pairs | eLife

TF-COMB – Discovering grammar of transcription factor binding sites - Computational and Structural Biotechnology Journal

Noncanonical binding of transcription factors: time to revisit specificity? | Molecular Biology of the Cell
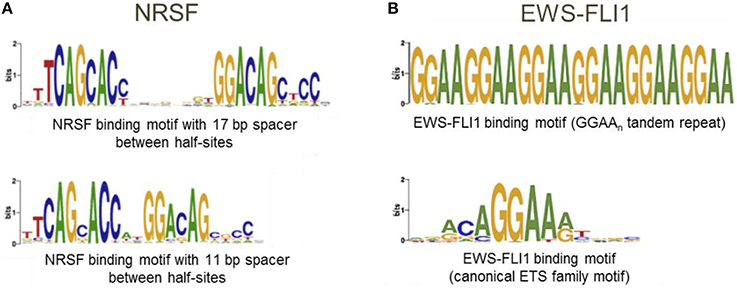
Frontiers | Analysis of Genomic Sequence Motifs for Deciphering Transcription Factor Binding and Transcriptional Regulation in Eukaryotic Cells

Curated collection of yeast transcription factor DNA binding specificity data reveals novel structural and gene regulatory insights | Genome Biology | Full Text

Predicting transcription factor binding sites using DNA shape features based on shared hybrid deep learning architecture: Molecular Therapy - Nucleic Acids

How motif environment influences transcription factor search dynamics: Finding a needle in a haystack - Dror - 2016 - BioEssays - Wiley Online Library

Impact of cancer mutational signatures on transcription factor motifs in the human genome | BMC Medical Genomics | Full Text

Discovering unknown human and mouse transcription factor binding sites and their characteristics from ChIP-seq data | PNAS

Reversible Disruption of Specific Transcription Factor-DNA Interactions Using CRISPR/Cas9 - ScienceDirect

Figure 1 from Subtypes of associated protein–DNA (Transcription Factor-Transcription Factor Binding Site) patterns | Semantic Scholar

Transcription factor (TF)-binding motif analysis. Examples of altered... | Download Scientific Diagram