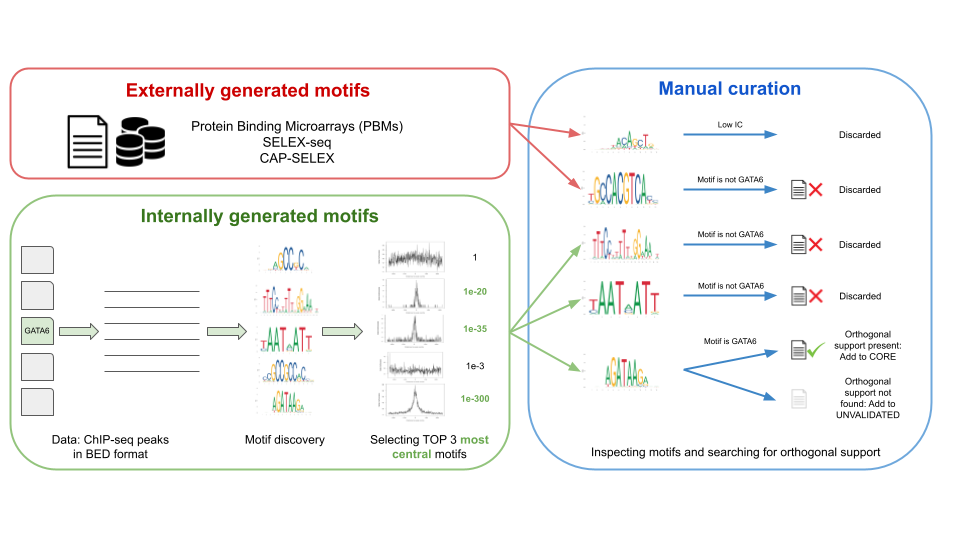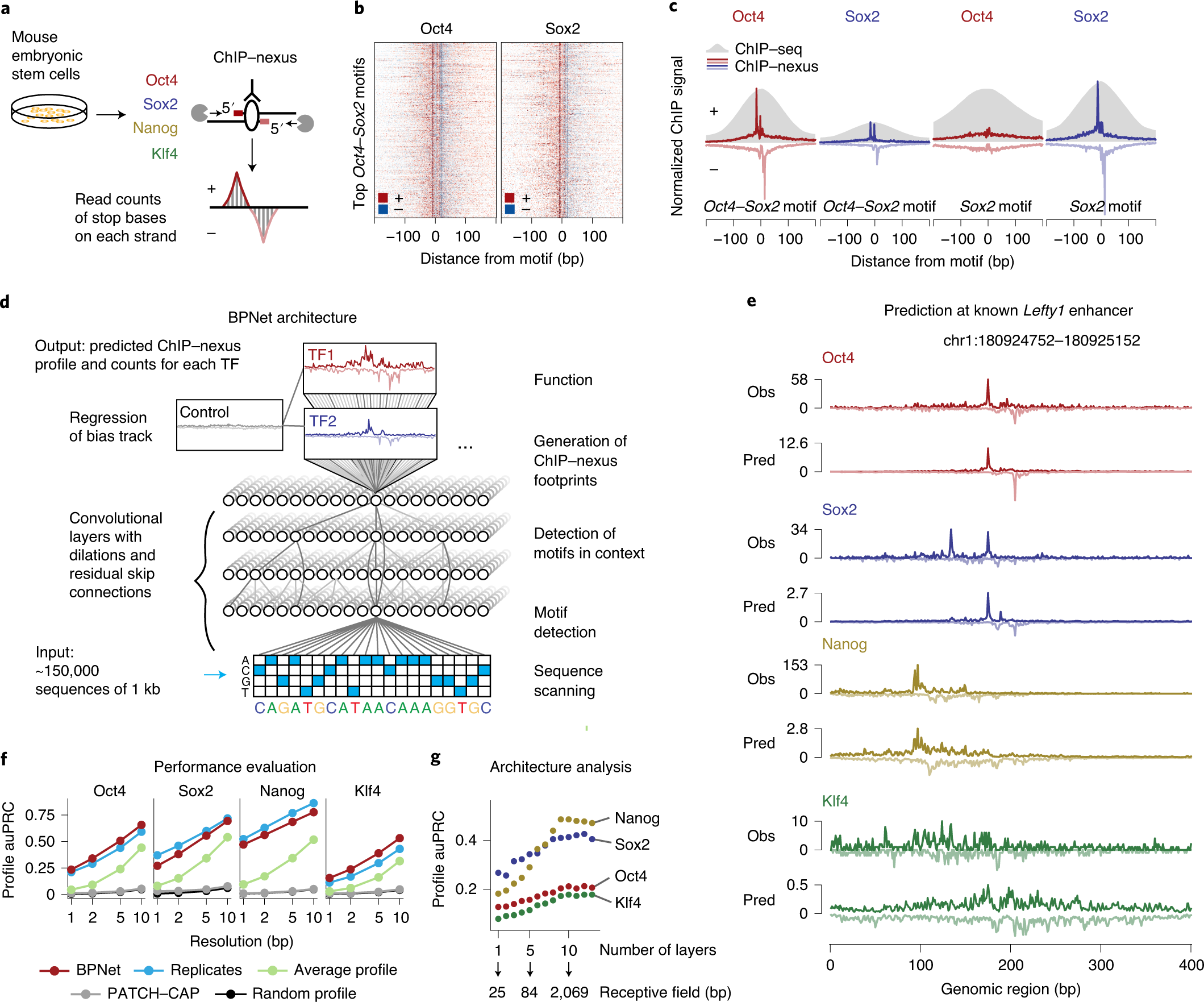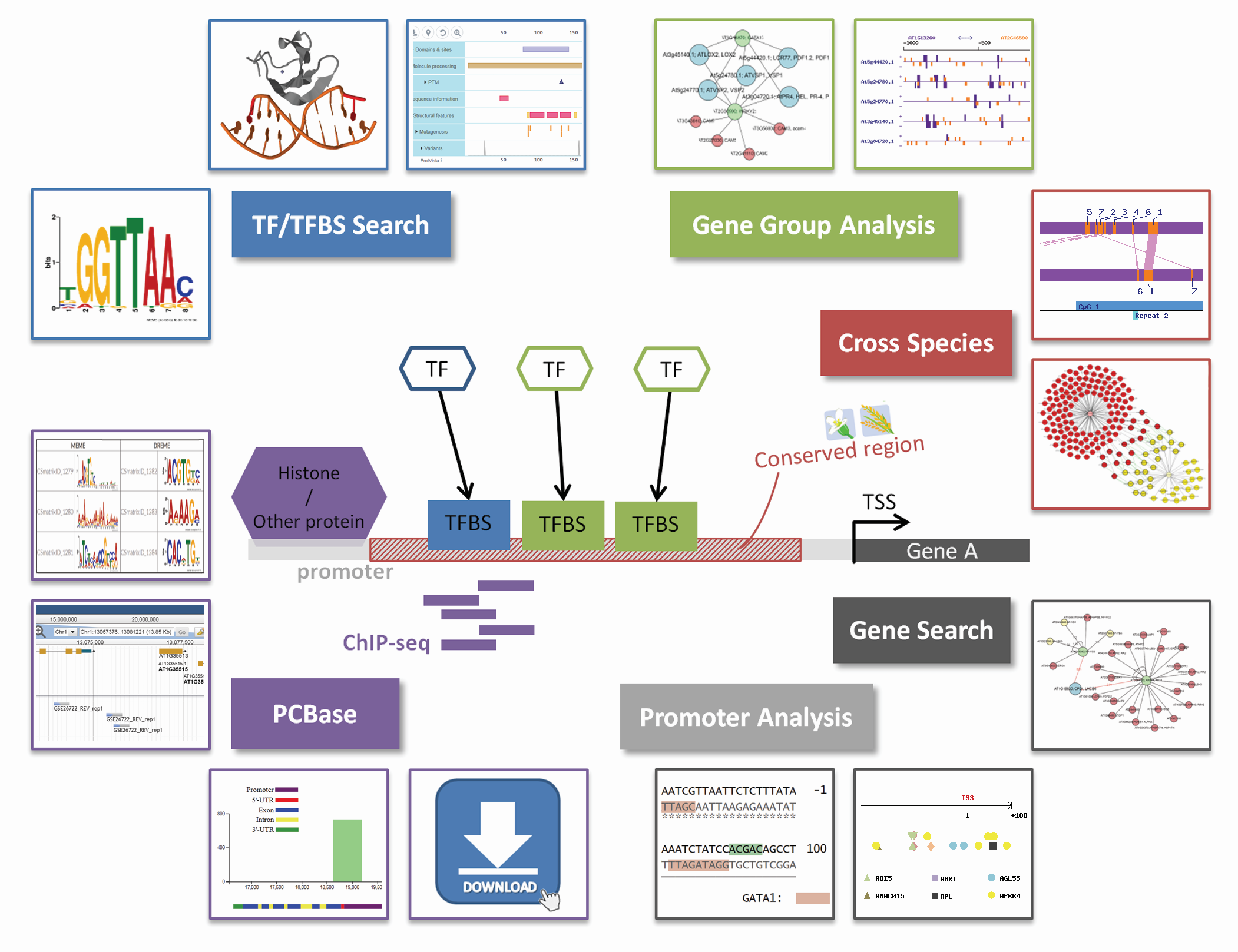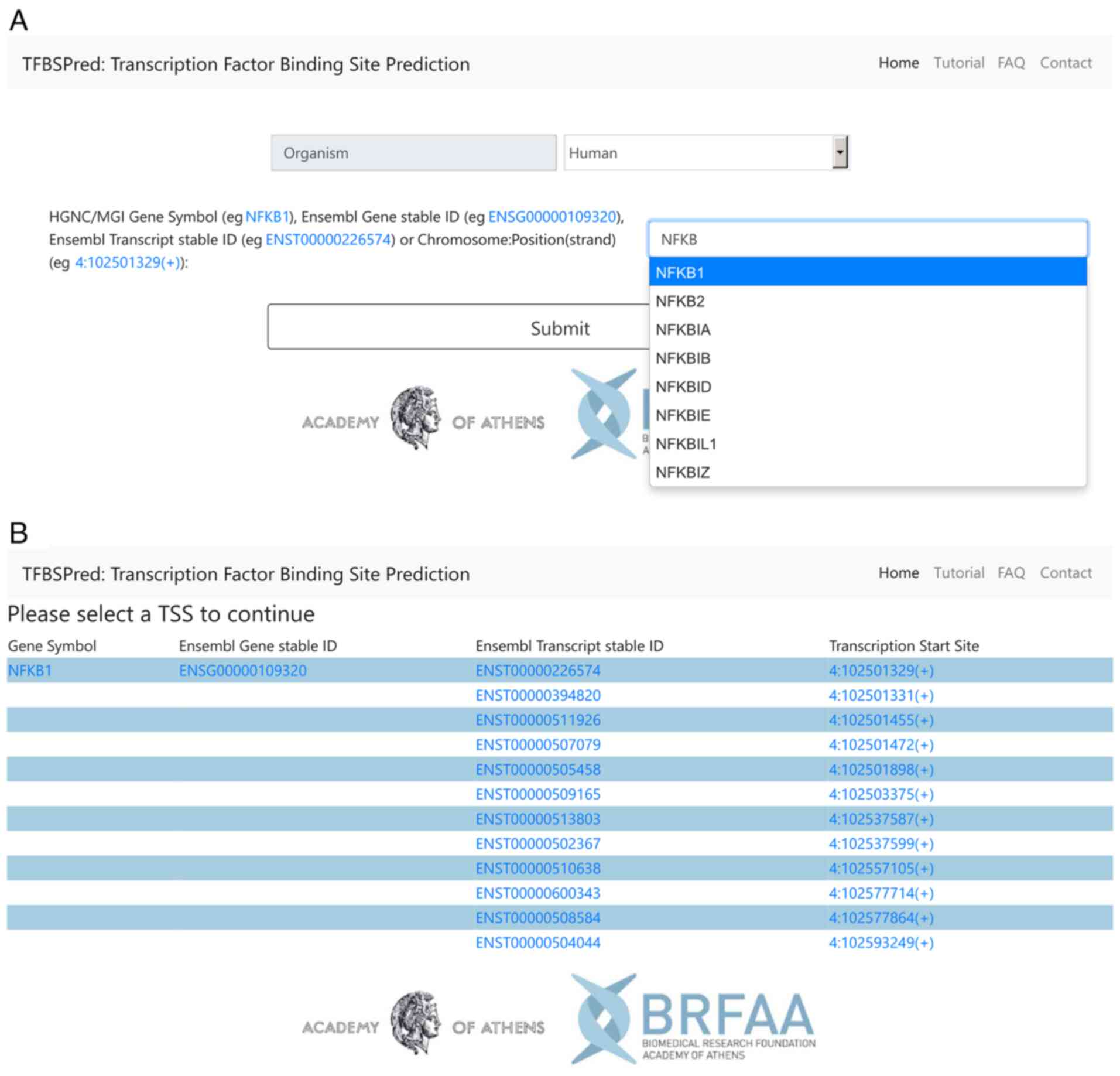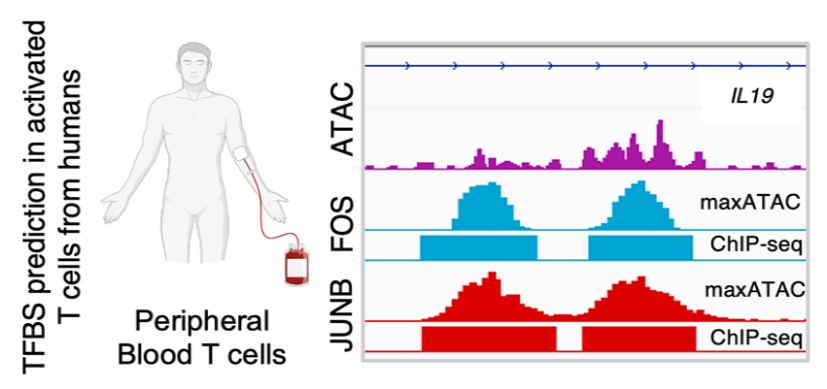
New Tool Provides Low-Cost, High-Quality Means to Map Transcription Factor Binding Genome-Wide - Research Horizons

Prediction of transcription factor binding sites and mutation analysis... | Download Scientific Diagram

DeepGRN: prediction of transcription factor binding site across cell-types using attention-based deep neural networks | BMC Bioinformatics | Full Text

Differences among the results of Grit and other prediction tools. A.... | Download Scientific Diagram
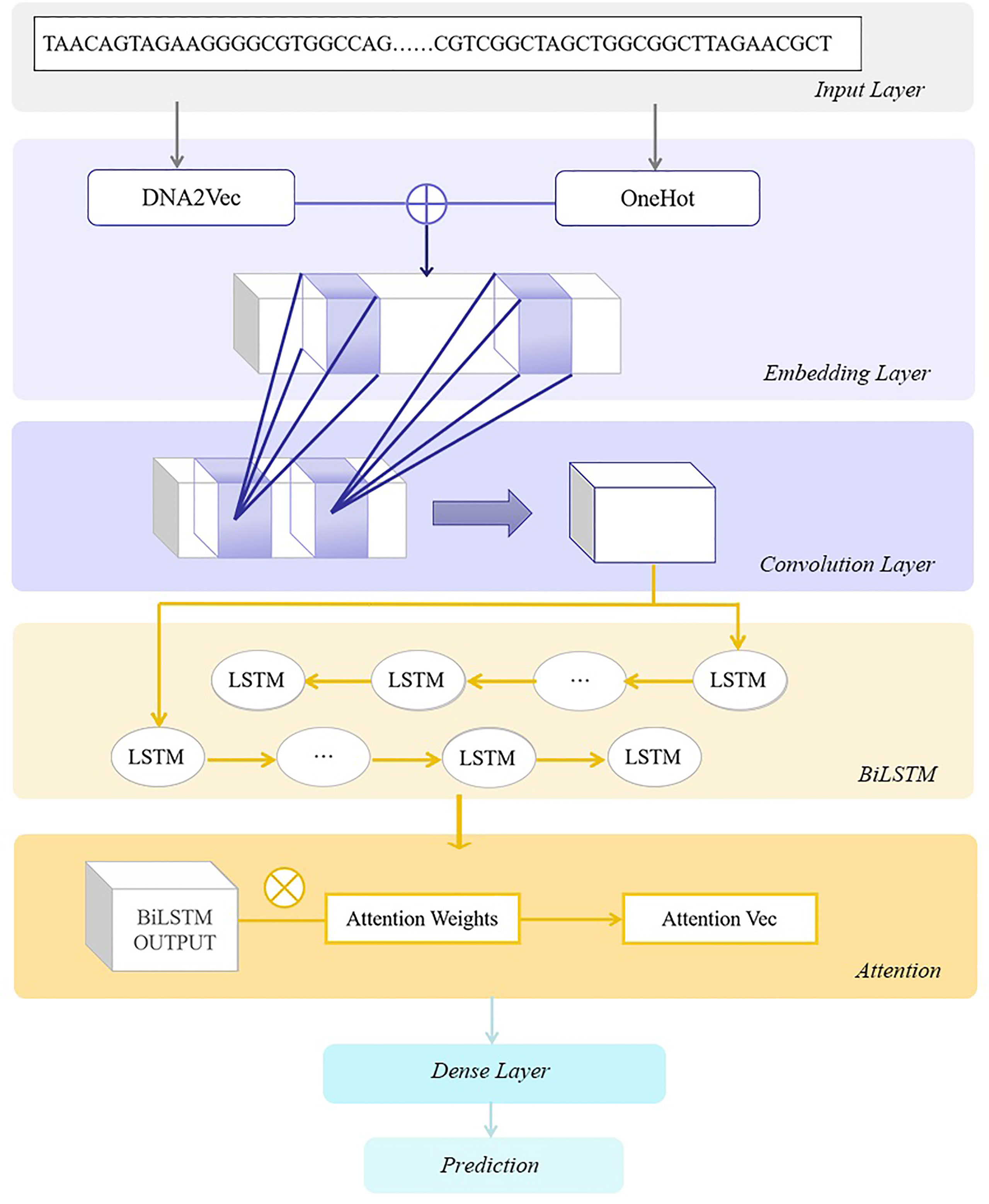
Frontiers | Prediction of Transcription Factor Binding Sites Using a Combined Deep Learning Approach
![PDF] TFBSTools: an R/bioconductor package for transcription factor binding site analysis | Semantic Scholar PDF] TFBSTools: an R/bioconductor package for transcription factor binding site analysis | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/7952b7c68b33972d5f8483cf7742efaed6396357/2-Figure1-1.png)
PDF] TFBSTools: an R/bioconductor package for transcription factor binding site analysis | Semantic Scholar

Predicting transcription factor binding sites using DNA shape features based on shared hybrid deep learning architecture: Molecular Therapy - Nucleic Acids