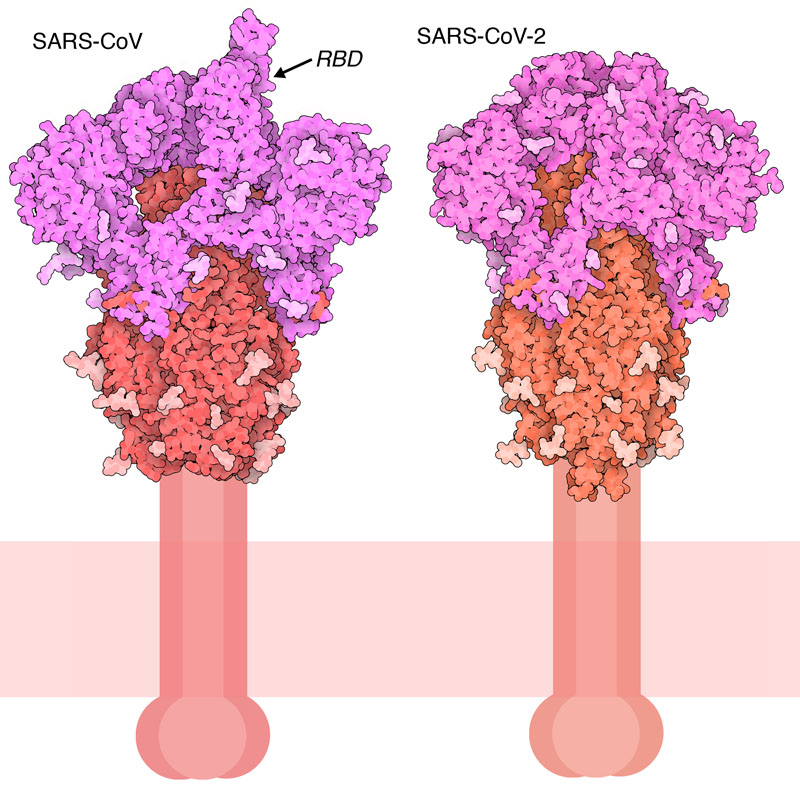
Do genetic polymorphisms in angiotensin converting enzyme 2 (ACE2) gene play a role in coronavirus disease 2019 (COVID-19)?

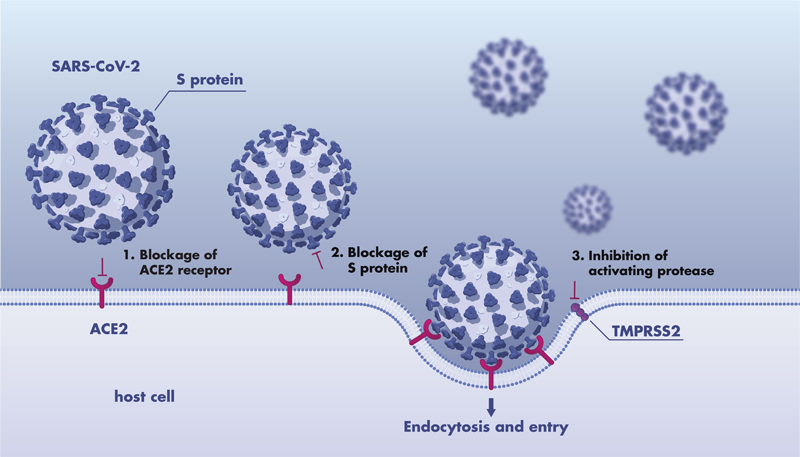
Synthetic α‐Helical Peptides as Potential Inhibitors of the ACE2 SARS‐CoV‐2 Interaction - Engelhardt - 2022 - ChemBioChem - Wiley Online Library

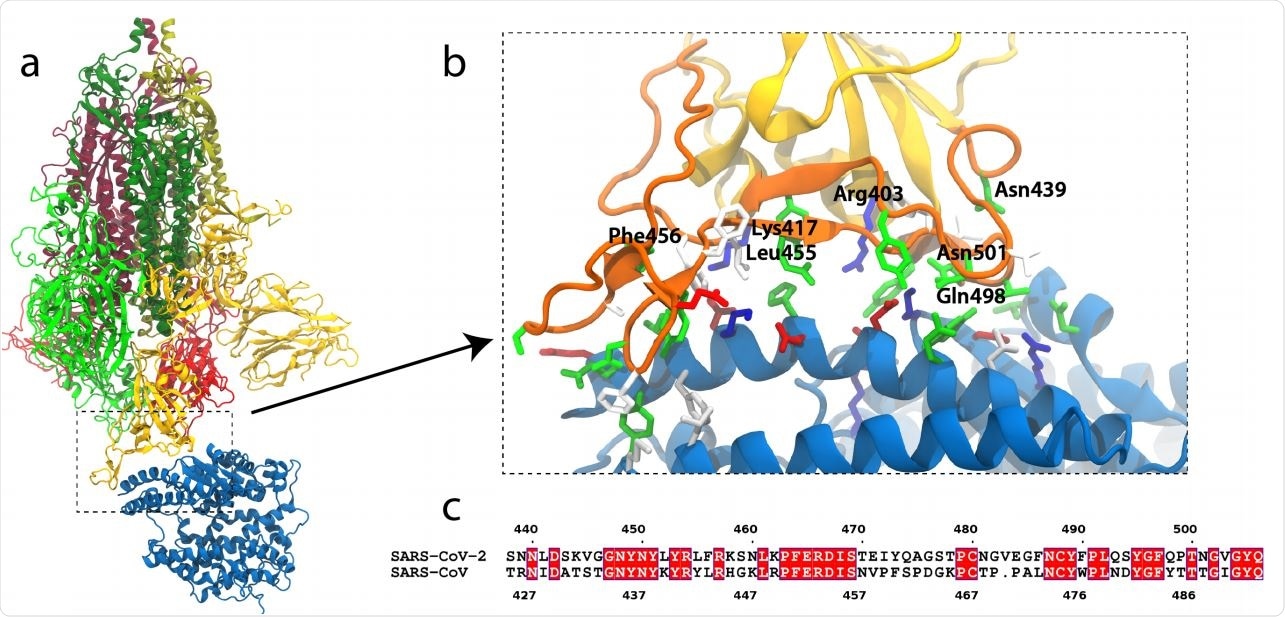
SARS-CoV-2 Omicron Variant: ACE2 Binding, Cryo-EM Structure of Spike Protein -ACE2 Complex and Antibody Evasion | bioRxiv
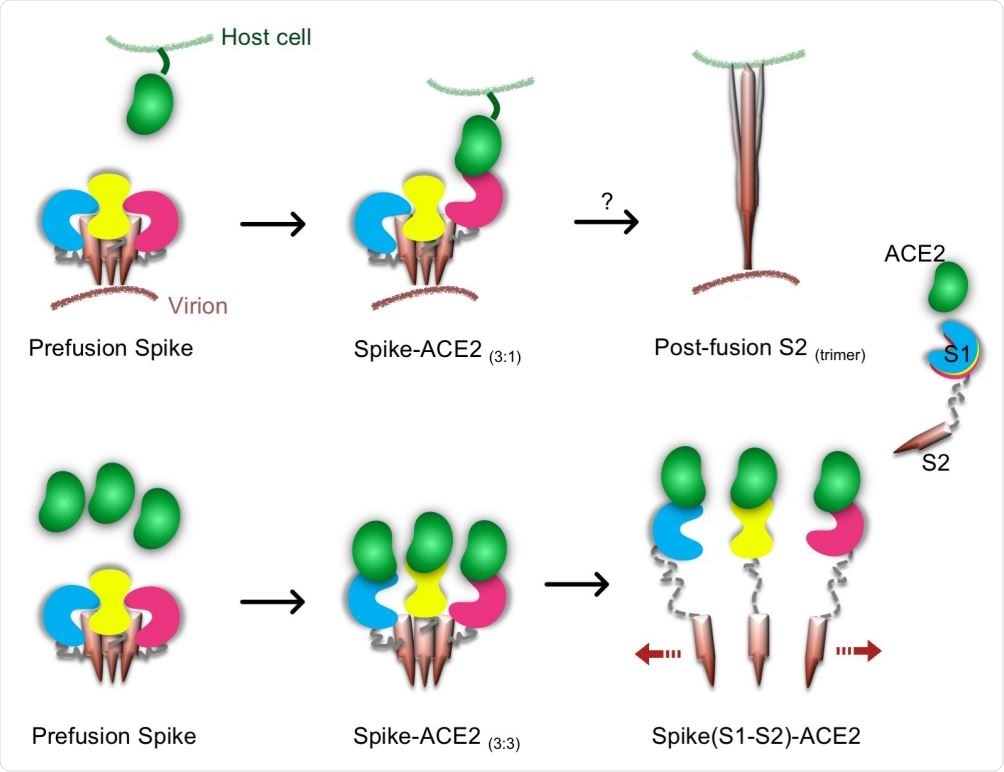
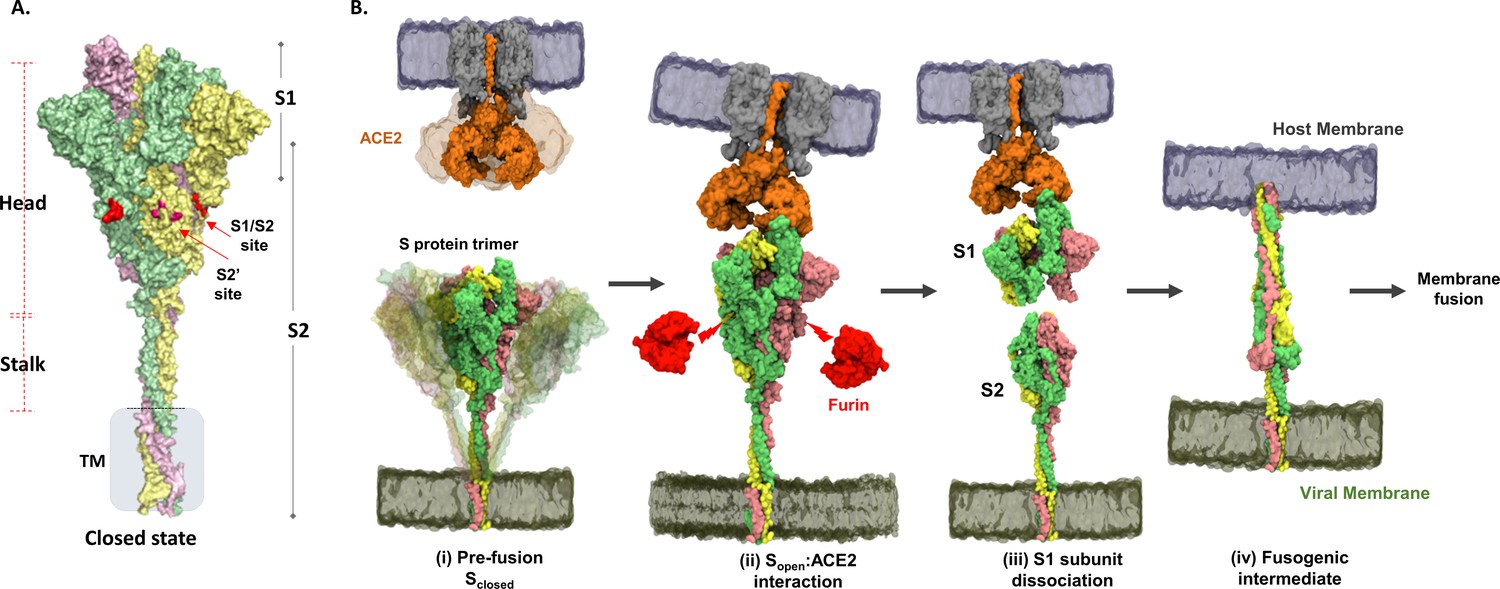
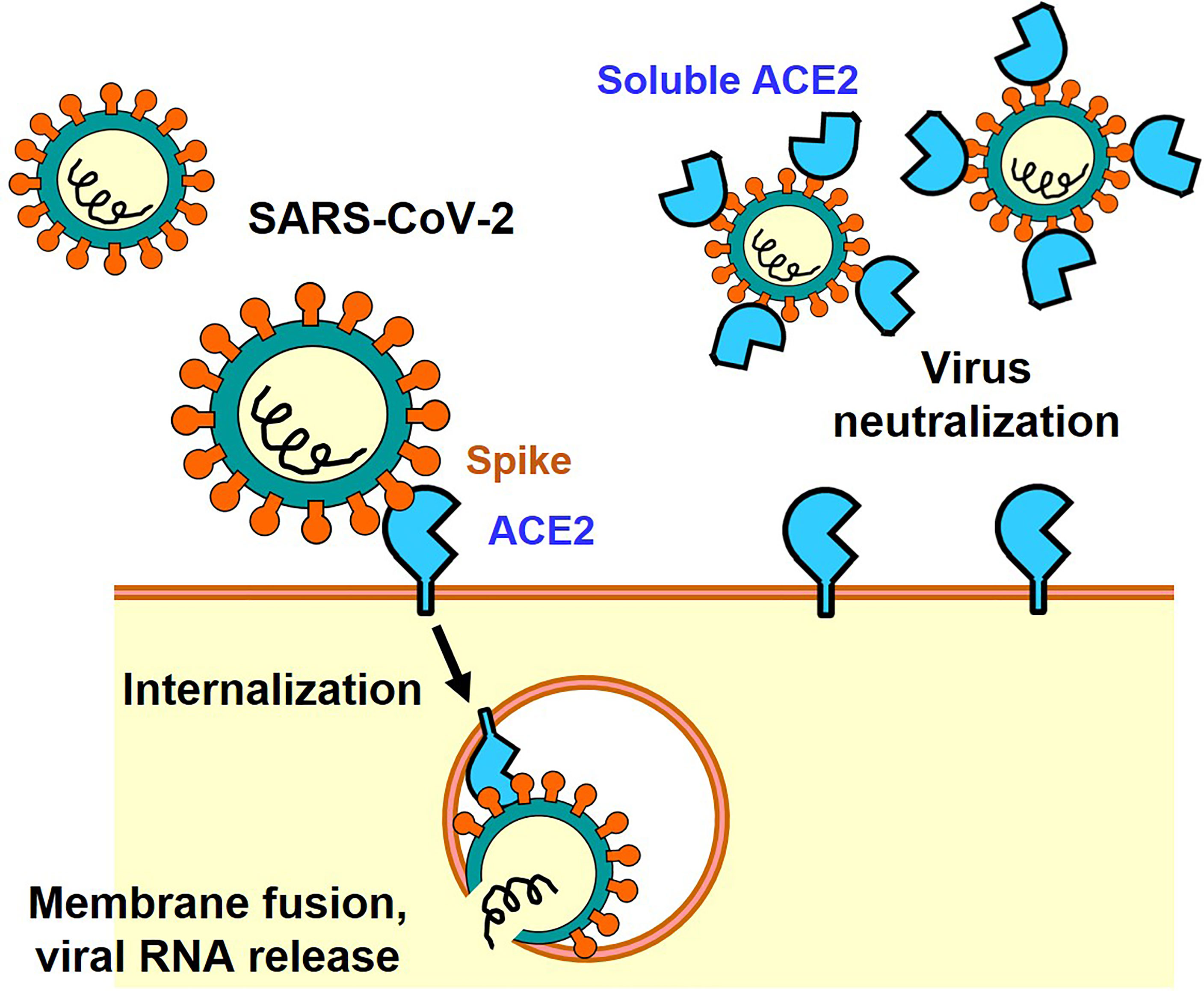
ACE2 binding causes dissociation of trimeric SARS-CoV-2 spike, may be used to inhibit further infection

SARS‐CoV‐2 first contact: Spike–ACE2 interactions in COVID‐19 - Nesci - 2021 - Chemical Biology & Drug Design - Wiley Online Library

Molecular interaction and inhibition of SARS-CoV-2 binding to the ACE2 receptor | Nature Communications
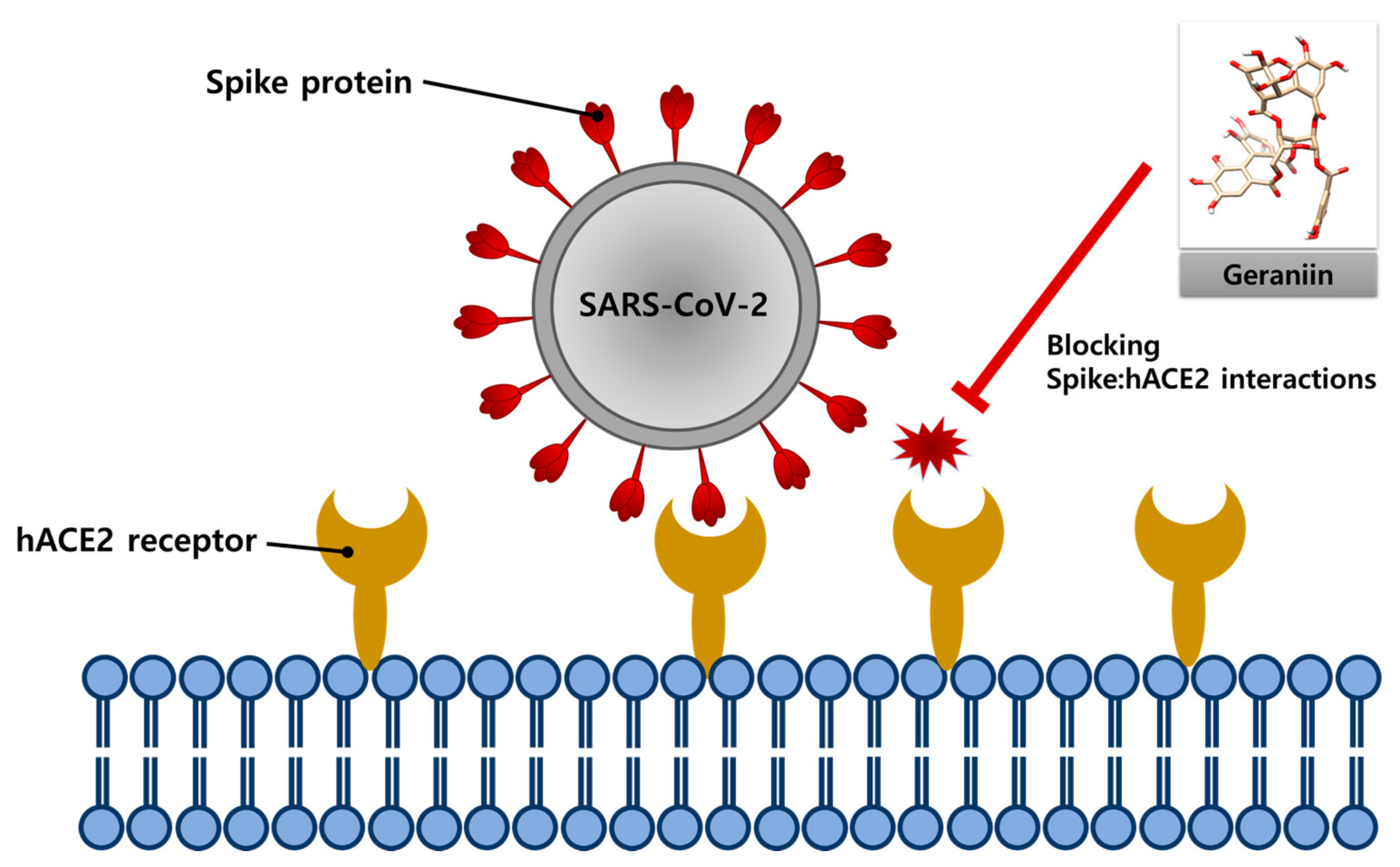
IJMS | Free Full-Text | Geraniin Inhibits the Entry of SARS-CoV-2 by Blocking the Interaction between Spike Protein RBD and Human ACE2 Receptor
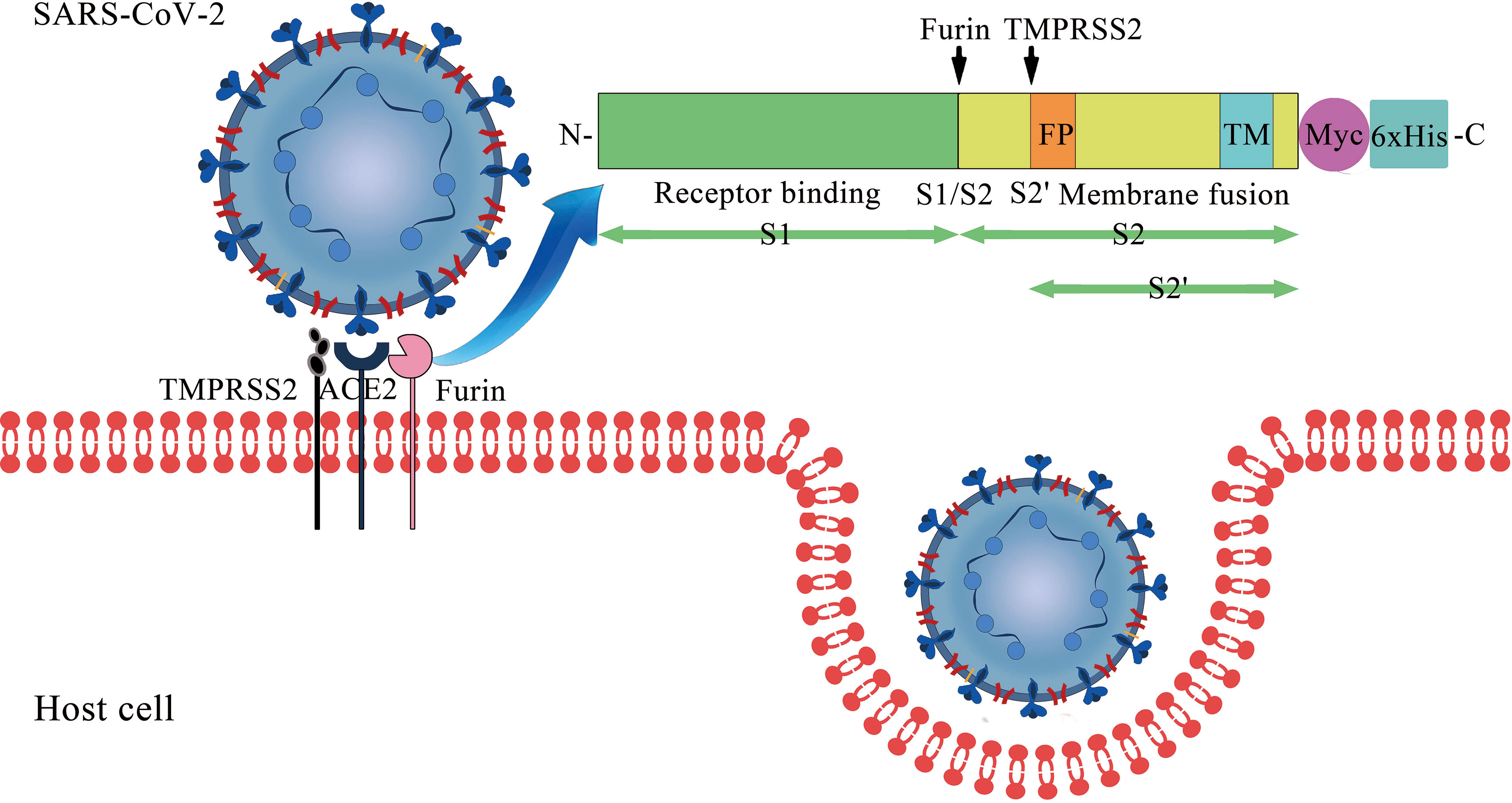
Frontiers | The Impact of ACE2 Polymorphisms on COVID-19 Disease: Susceptibility, Severity, and Therapy

Cryo-EM Structures of SARS-CoV-2 Spike without and with ACE2 Reveal a pH-Dependent Switch to Mediate Endosomal Positioning of Receptor-Binding Domains - ScienceDirect
Development and simulation of fully glycosylated molecular models of ACE2-Fc fusion proteins and their interaction with the SARS-CoV-2 spike protein binding domain | PLOS ONE
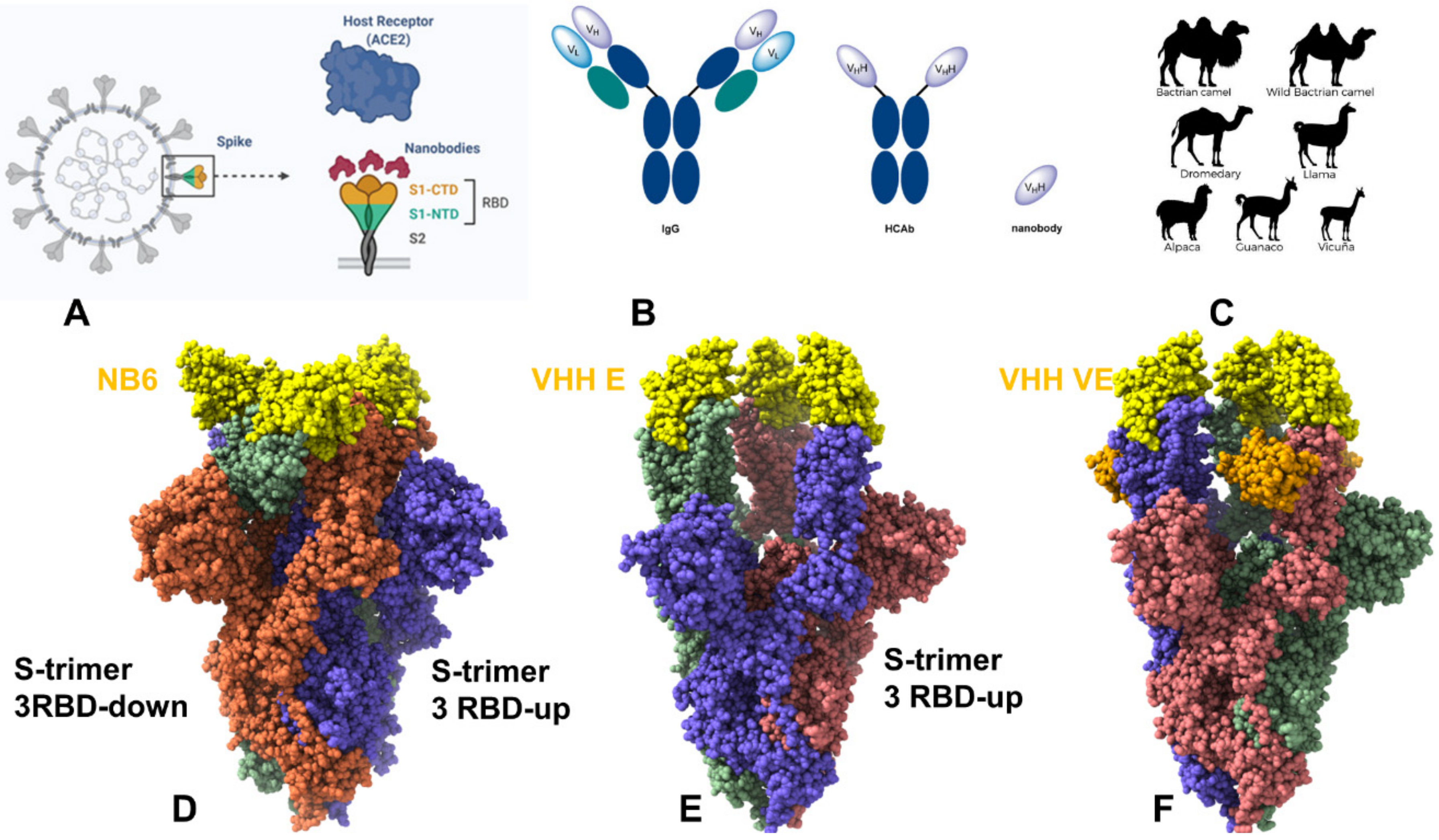
IJMS | Free Full-Text | Structural and Computational Studies of the SARS-CoV-2 Spike Protein Binding Mechanisms with Nanobodies: From Structure and Dynamics to Avidity-Driven Nanobody Engineering

Engaging the spikes: heparan sulfate facilitates SARS-CoV-2 spike protein binding to ACE2 and potentiates viral infection | Signal Transduction and Targeted Therapy

Computational biophysical characterization of the SARS-CoV-2 spike protein binding with the ACE2 receptor and implications for infectivity - Computational and Structural Biotechnology Journal

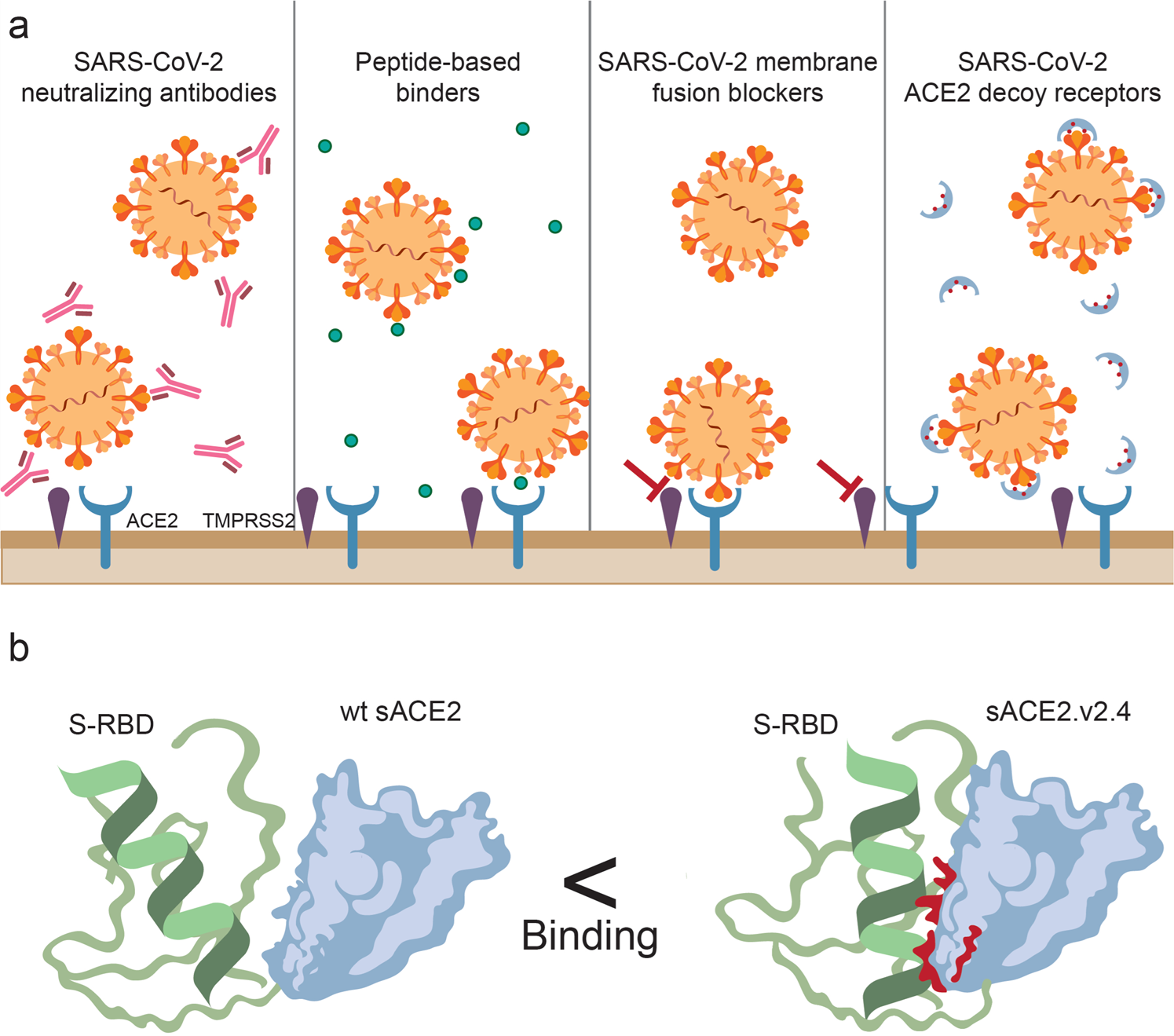
An Update on Novel Soluble ACE2 Therapeutics to Treat SARS-CoV-2: Insights from a Preclinical Study in: Kidney News Volume 13 Issue 3 (2021)
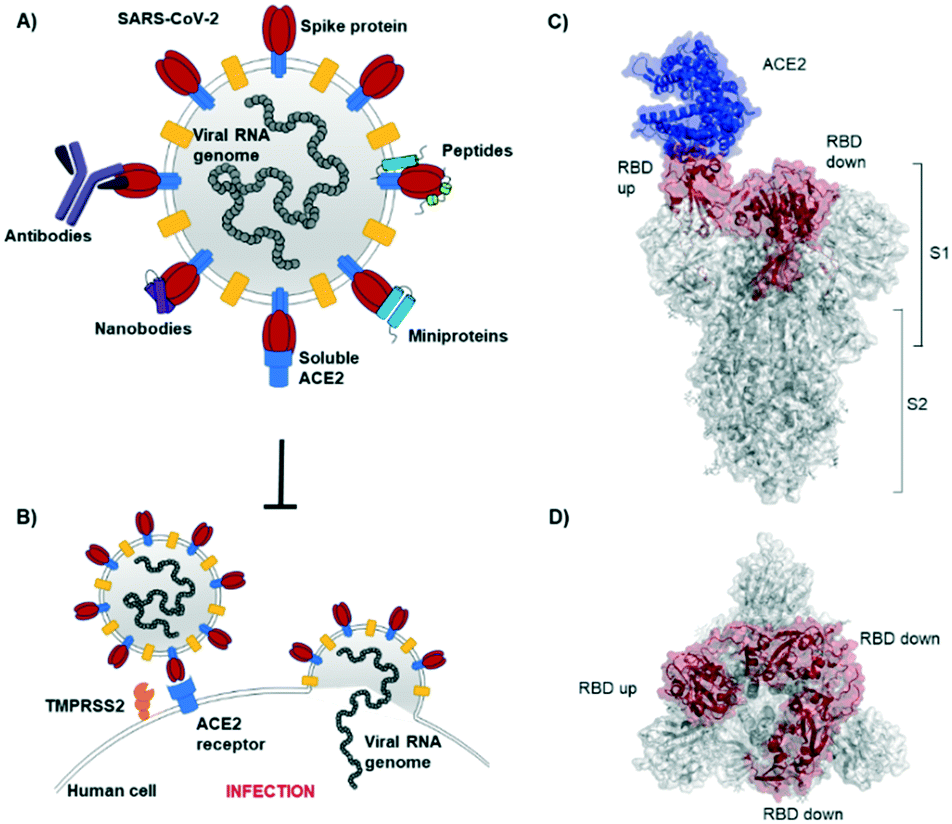
Targeting the SARS-CoV-2-spike protein: from antibodies to miniproteins and peptides - RSC Medicinal Chemistry (RSC Publishing) DOI:10.1039/D0MD00385A

Researchers discover potential therapeutic targets on SARS-CoV-2 Spike protein | Penn State University

Peptide modelling and screening against human ACE2 and spike glycoprotein RBD of SARS-CoV-2 | In Silico Pharmacology