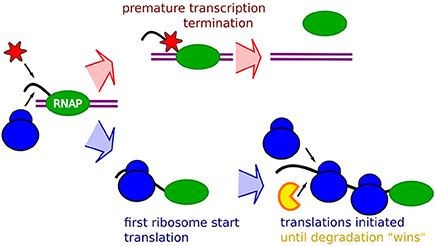
Frontiers | Occlusion of the Ribosome Binding Site Connects the Translational Initiation Frequency, mRNA Stability and Premature Transcription Termination

A set of synthetic versatile genetic control elements for the efficient expression of genes in Actinobacteria | Scientific Reports

Characterization and implications of prokaryotic ribosome-binding sites across species | Systems Microbiology and Biomanufacturing
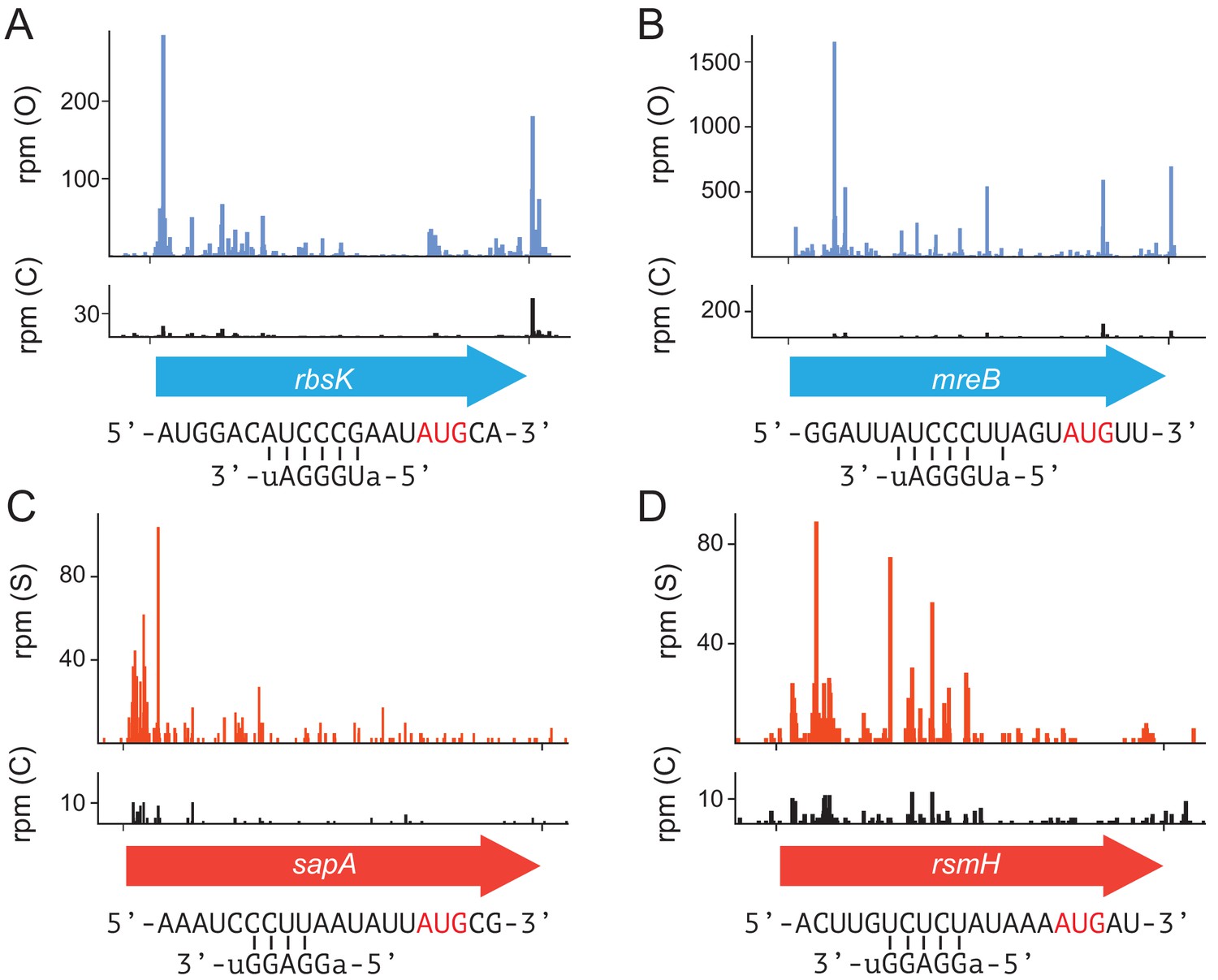
Translational initiation in E. coli occurs at the correct sites genome-wide in the absence of mRNA-rRNA base-pairing | eLife
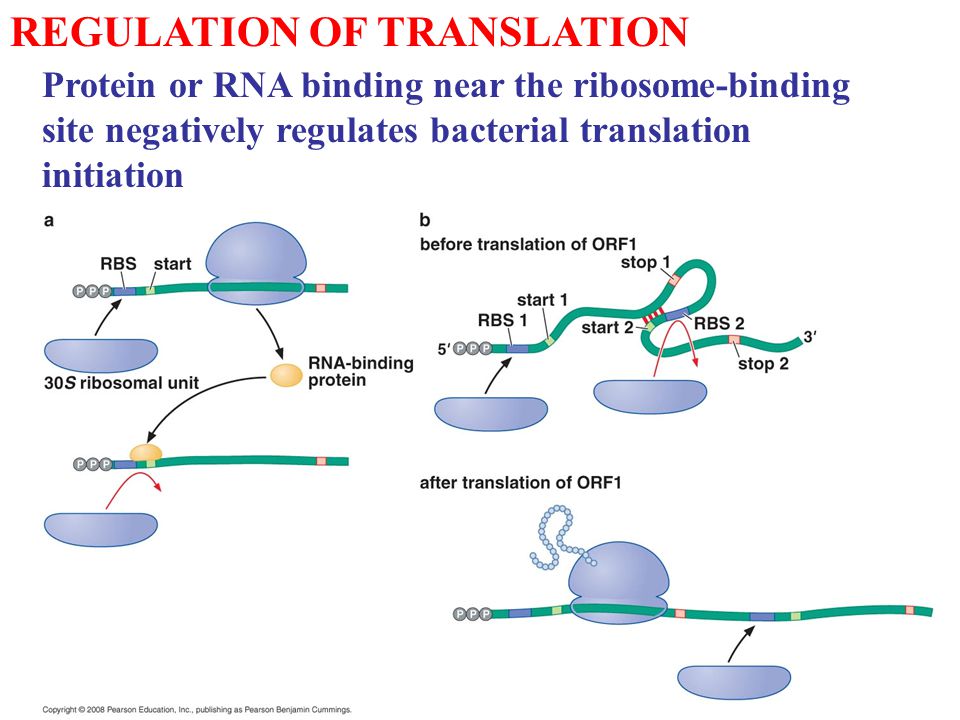
REGULATION OF TRANSLATION Protein or RNA binding near the ribosome-binding site negatively regulates bacterial translation initiation. - ppt download

Deciphering the Rules of Ribosome Binding Site Differentiation in Context Dependence | ACS Synthetic Biology

rClone: A Synthetic Biology Tool That Enables the Research of Bacterial Translation — Journal of Young Investigators
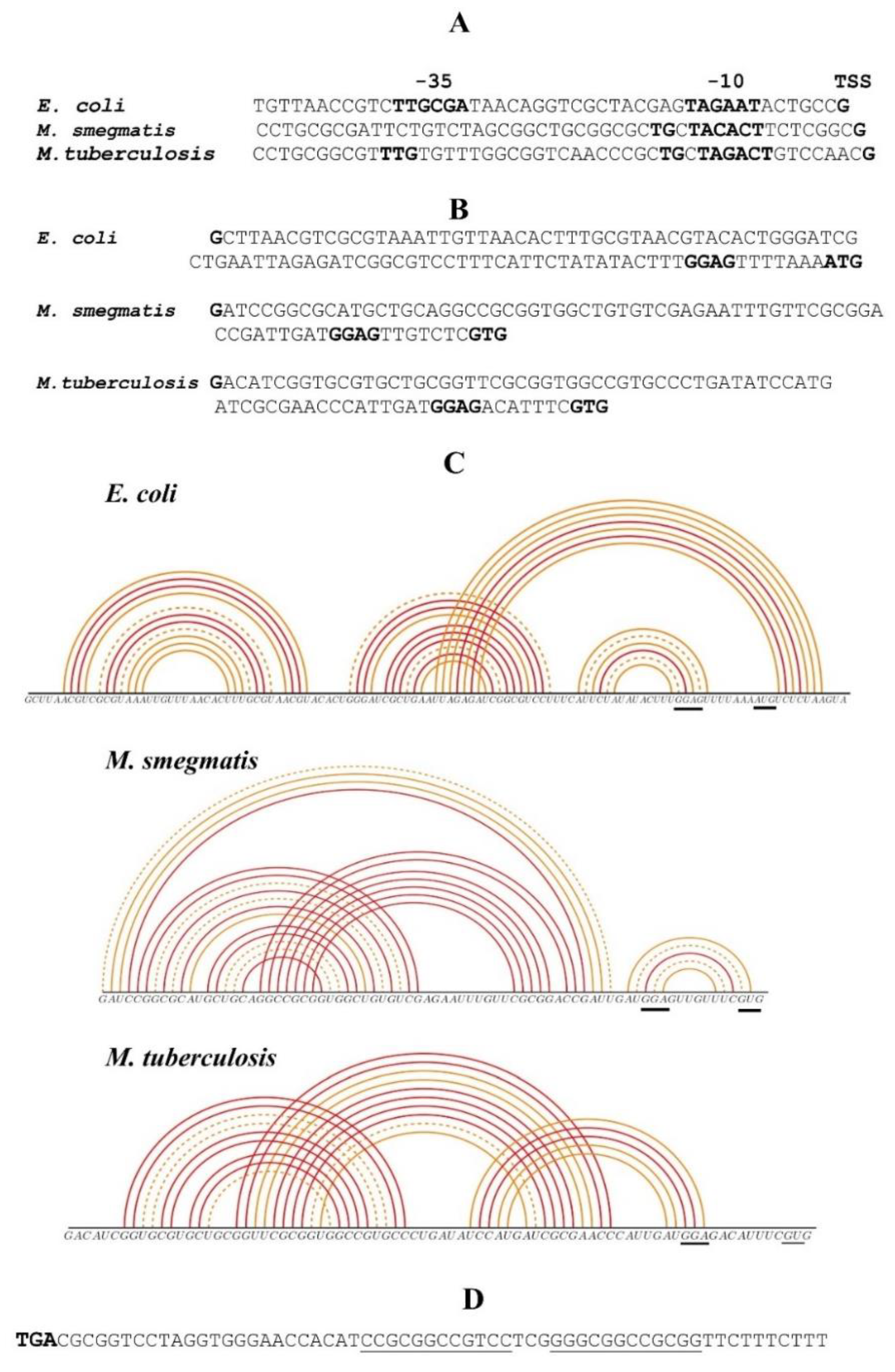
IJMS | Free Full-Text | Regulation of Ribosomal Protein Synthesis in Mycobacteria: The Autogenous Control of rpsO

The length of ribosomal binding site spacer sequence controls the production yield for intracellular and secreted proteins by Bacillus subtilis | Microbial Cell Factories | Full Text

Sequence analysis of the nic promoters. (a) The ribosome binding sites... | Download Scientific Diagram

Ribosomal binding site sequences and promoters for expressing glutamate decarboxylase and producing γ-aminobutyrate in Corynebacterium glutamicum | AMB Express | Full Text

Ribosome binding site libraries and pathway modules for shikimic acid synthesis with Corynebacterium glutamicum | Microbial Cell Factories | Full Text

The length of ribosomal binding site spacer sequence controls the production yield for intracellular and secreted proteins by Bacillus subtilis | Microbial Cell Factories | Full Text
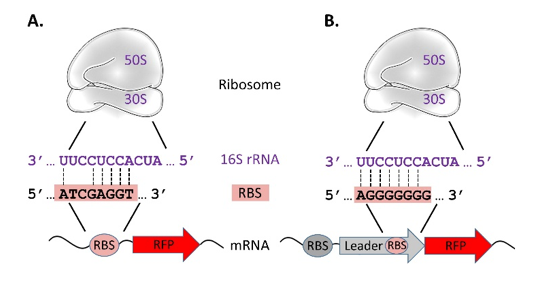
rClone: A Synthetic Biology Tool That Enables the Research of Bacterial Translation — Journal of Young Investigators

Determination of the ribosome-binding sequence and spacer length between binding site and initiation codon for efficient protein expression in Bifidobacterium longum 105-A - ScienceDirect


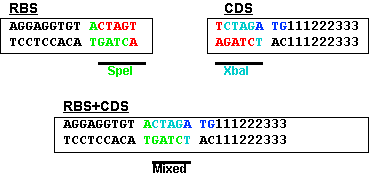


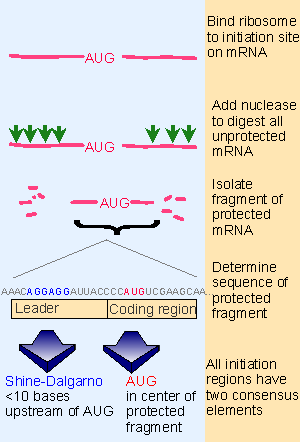

![PDF] Classification of Escherichia coli K-12 ribosome binding sites | Semantic Scholar PDF] Classification of Escherichia coli K-12 ribosome binding sites | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/4af616123bce83b1f5dff449961c1df697c8d771/2-Figure2-1.png)