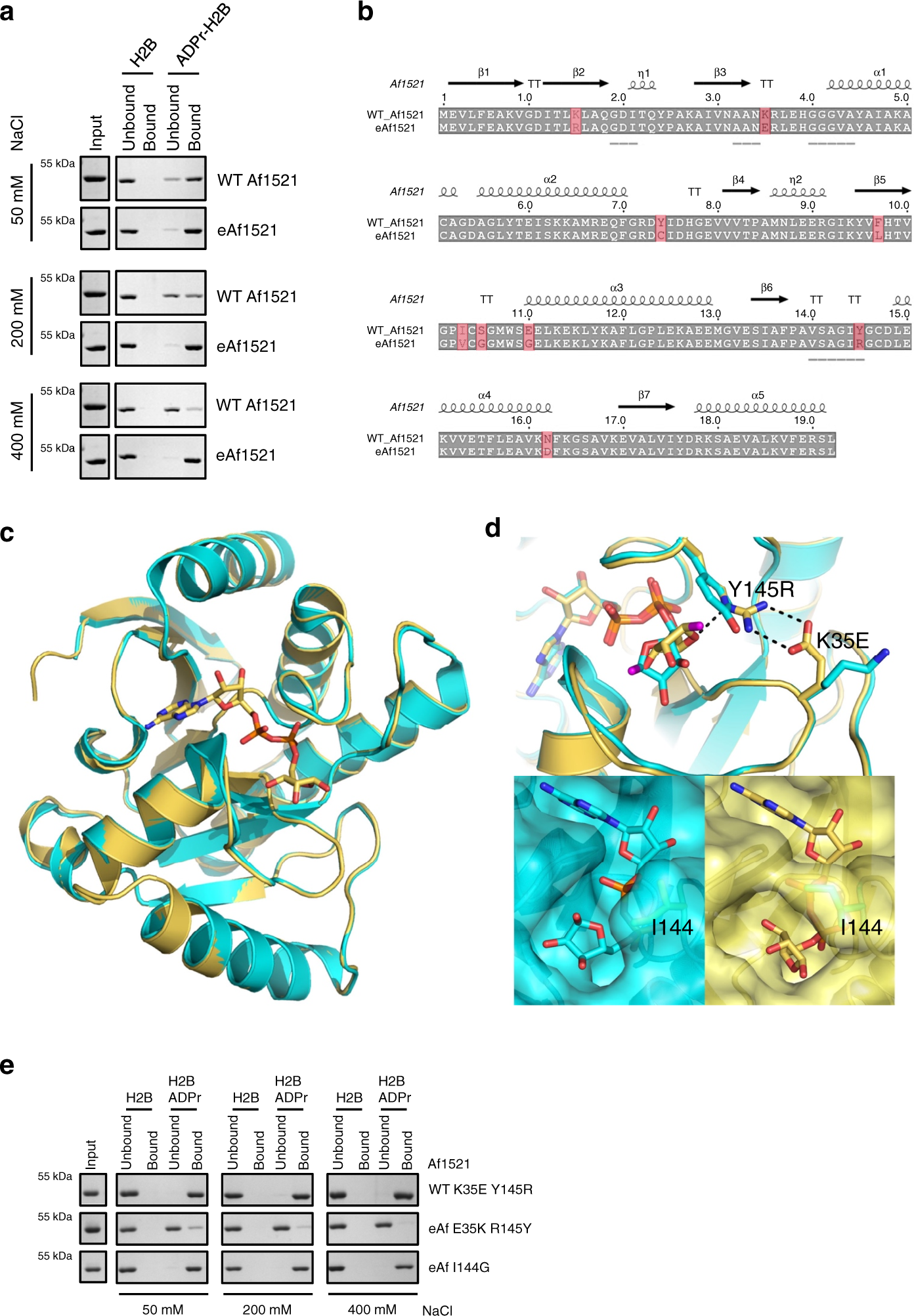
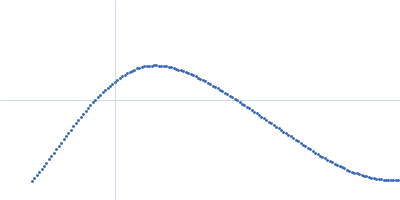
Generation and Characterization of Recombinant Antibody-like ADP-Ribose Binding Proteins | Biochemistry

The role of solute binding proteins in signal transduction - Computational and Structural Biotechnology Journal

Generation and Characterization of Recombinant Antibody-like ADP-Ribose Binding Proteins | Biochemistry

Structures of the G134R mutant of ribose binding protein (RBPG134R) in... | Download Scientific Diagram

Procedure for morphing and recapitulating an intermediate, with Ribose... | Download Scientific Diagram
Probing protein-protein interactions. The ribose-binding protein in bacterial transport and chemotaxis.
Selective monitoring of the protein-free ADP-ribose released by ADP-ribosylation reversal enzymes | PLOS ONE

Unbound: In the absence of ribose, the ribose-binding protein (RBP) is... | Download Scientific Diagram

Identifying Poly(ADP-ribose)-Binding Proteins with Photoaffinity-Based Proteomics | Journal of the American Chemical Society

Computational redesign of the Escherichia coli ribose-binding protein ligand binding pocket for 1,3-cyclohexanediol and cyclohexanol | Scientific Reports
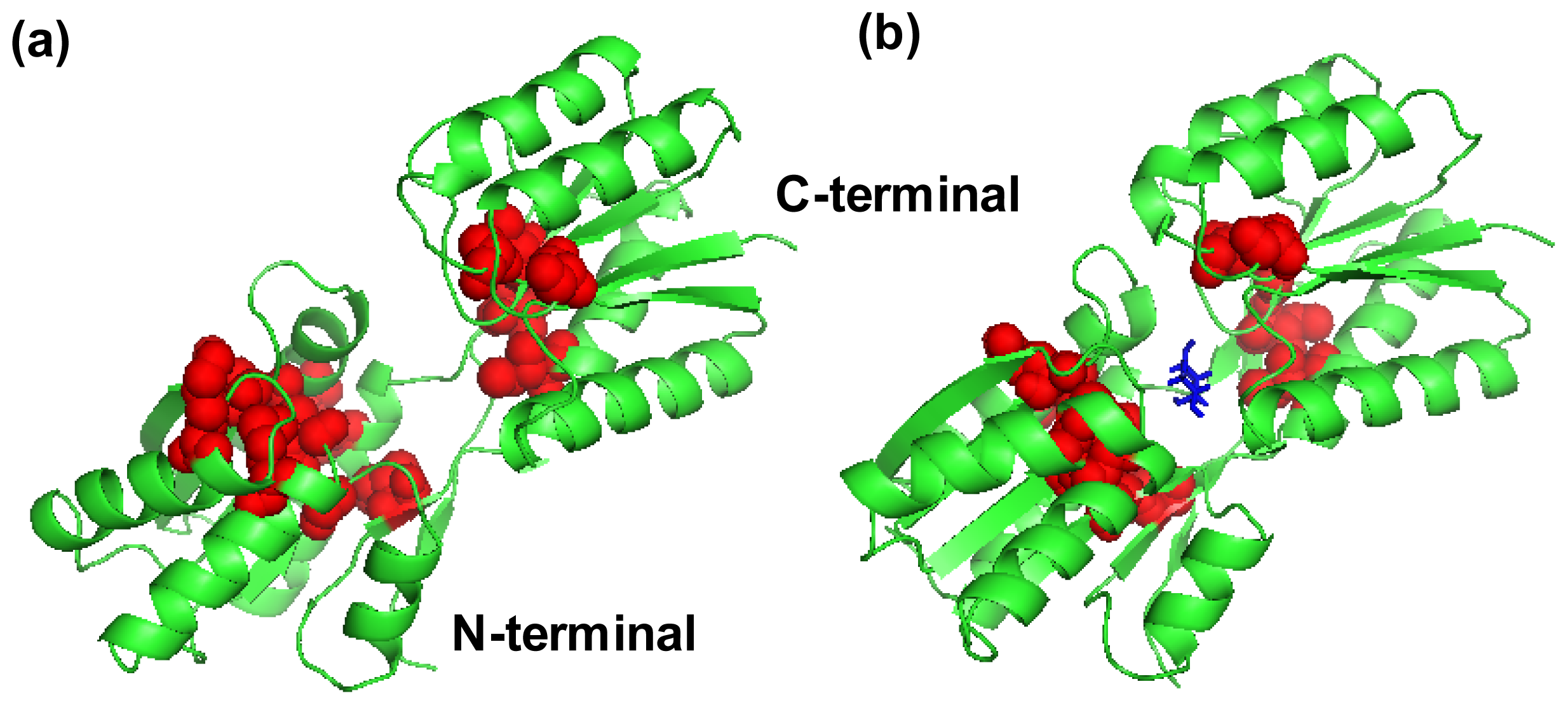
IJMS | Free Full-Text | Analysis of Conformational Motions and Residue Fluctuations for Escherichia coli Ribose-Binding Protein Revealed with Elastic Network Models

IJMS | Free Full-Text | Poly(ADP-Ribose) Polymerases in Plants and Their Human Counterparts: Parallels and Peculiarities

Periplasmic-binding protein-based biosensors and bioanalytical assay platforms: Advances, considerations, and strategies for optimal utility - ScienceDirect
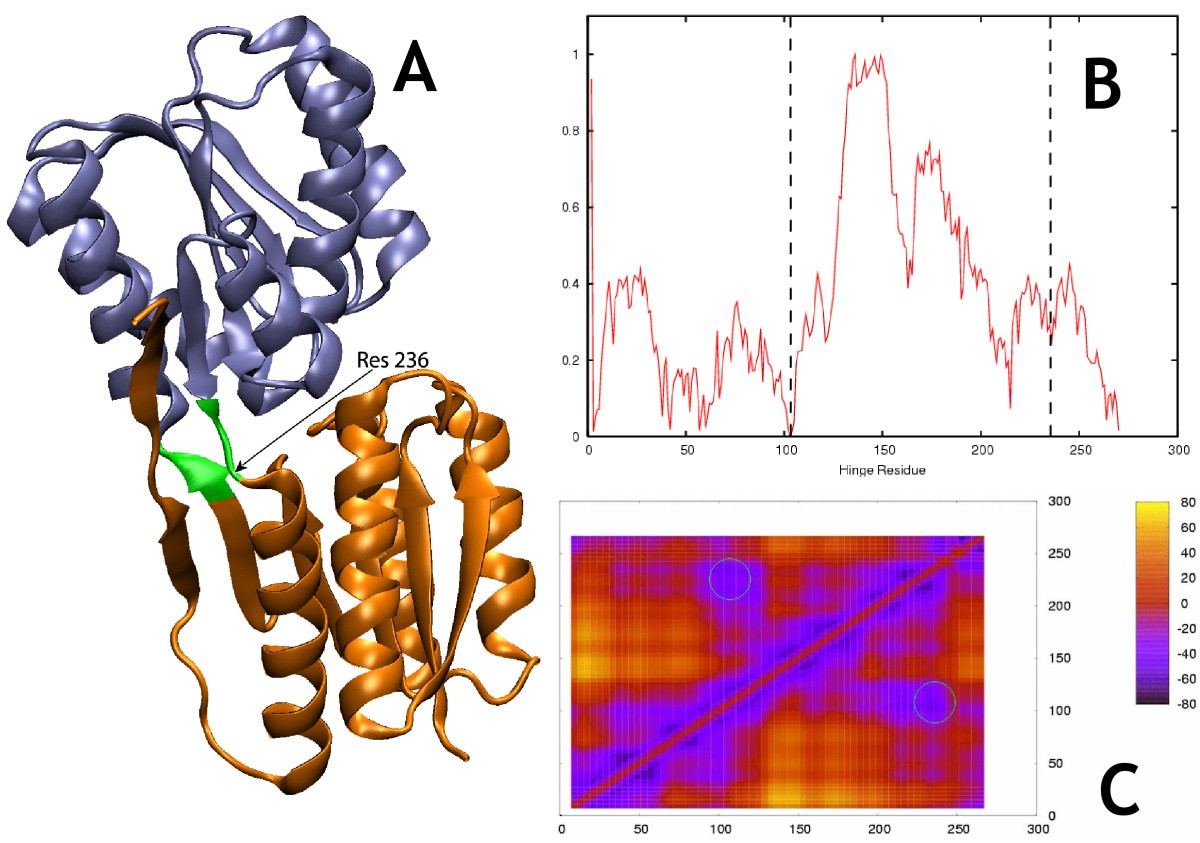
FlexOracle: predicting flexible hinges by identification of stable domains | BMC Bioinformatics | Full Text
Probing protein-protein interactions. The ribose-binding protein in bacterial transport and chemotaxis.

Computational redesign of the Escherichia coli ribose-binding protein ligand binding pocket for 1,3-cyclohexanediol and cyclohexanol | Scientific Reports