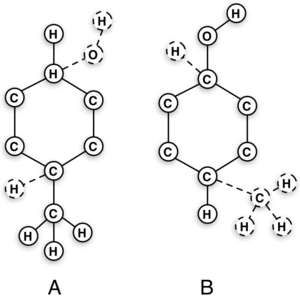
7α-18F-Fluoromethyl-Dihydrotestosterone and 7α-18F-Fluoromethyl-Nortestosterone: Ligands to Determine the Role of Sex Hormone–Binding Globulin for Steroidal Radiopharmaceuticals | Journal of Nuclear Medicine

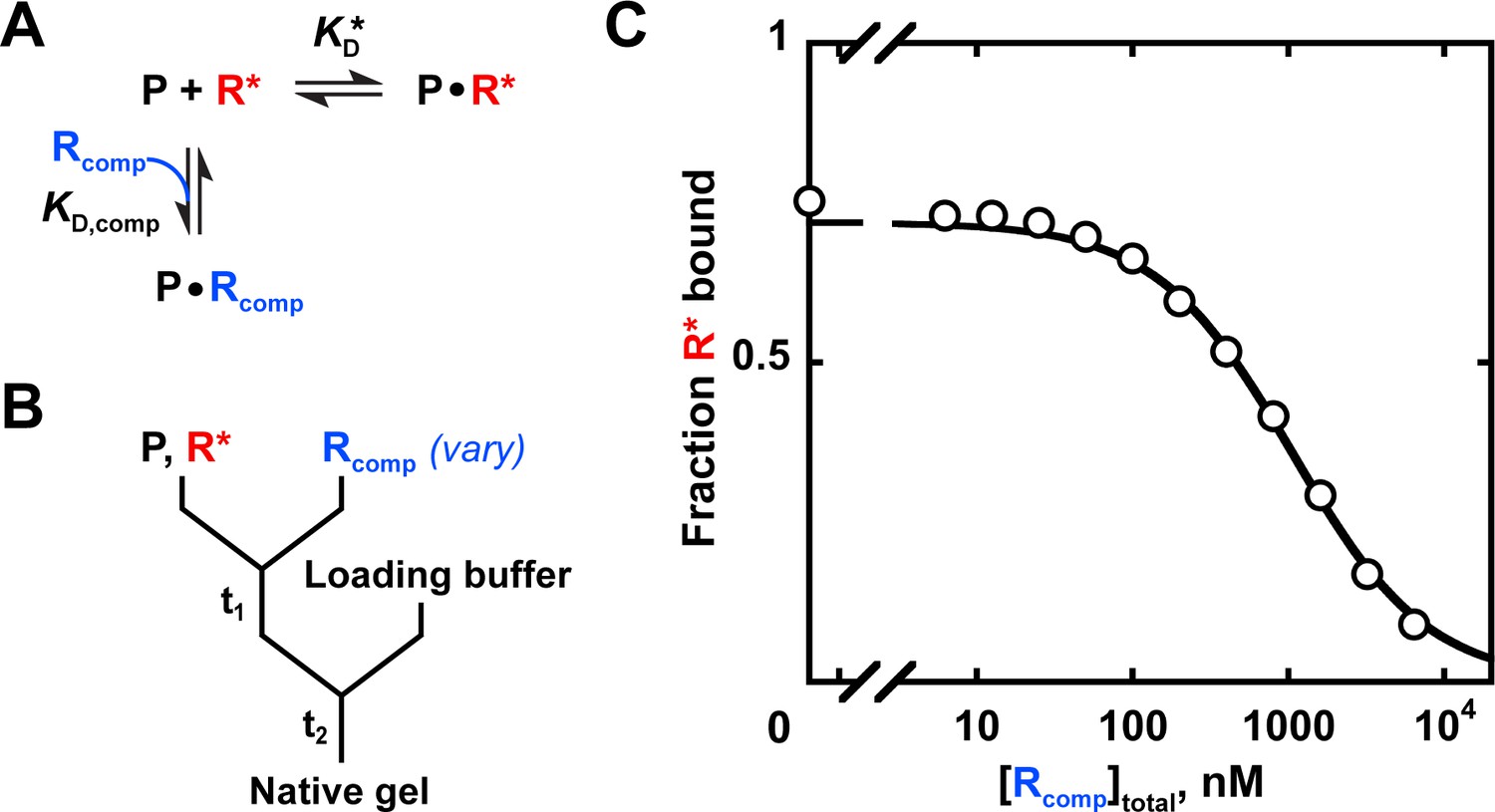
On the binding affinity of macromolecular interactions: daring to ask why proteins interact | Journal of The Royal Society Interface

PBCNet:Computing Relative Binding Affinity of Ligands to a Receptor Based on a Pairwise Binding Comparison Network for Lead Optimization | Theoretical and Computational Chemistry | ChemRxiv | Cambridge Open Engage

Predicting the Relative Binding Affinity for Reversible Covalent Inhibitors by Free Energy Perturbation Calculations | Journal of Chemical Information and Modeling

Relative Binding Free Energy Calculations in Drug Discovery: Recent Advances and Practical Considerations | Journal of Chemical Information and Modeling

GRAM: A True Null Model for Relative Binding Affinity Predictions | Theoretical and Computational Chemistry | ChemRxiv | Cambridge Open Engage

On the binding affinity of macromolecular interactions: daring to ask why proteins interact | Journal of The Royal Society Interface

Production and characterization of human anti-V3 monoclonal antibodies from the cells of HIV-1 infected Indian donors | Virology Journal | Full Text
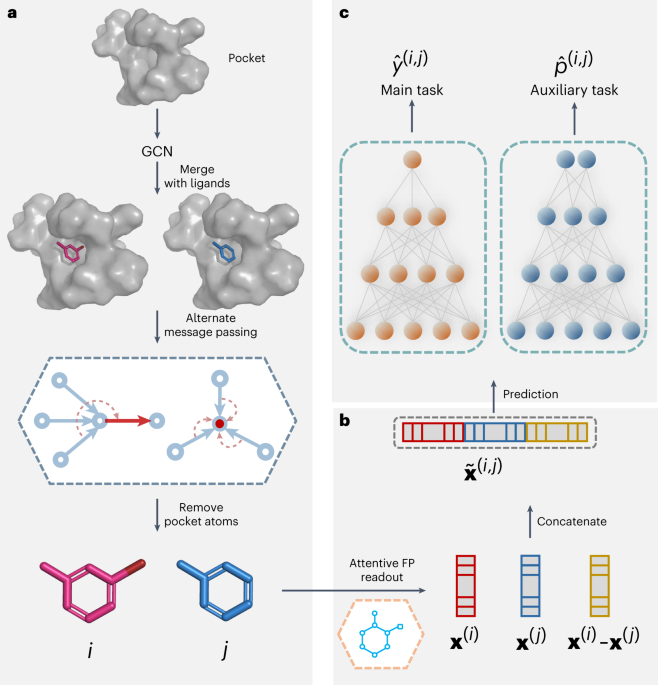
Computing the relative binding affinity of ligands based on a pairwise binding comparison network | Nature Computational Science
![PDF] The estrogen receptor relative binding affinities of 188 natural and xenochemicals: structural diversity of ligands. | Semantic Scholar PDF] The estrogen receptor relative binding affinities of 188 natural and xenochemicals: structural diversity of ligands. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/a4bc89a97a0457e76e7a503a7b14b8561bd870fb/5-Table3-1.png)
PDF] The estrogen receptor relative binding affinities of 188 natural and xenochemicals: structural diversity of ligands. | Semantic Scholar
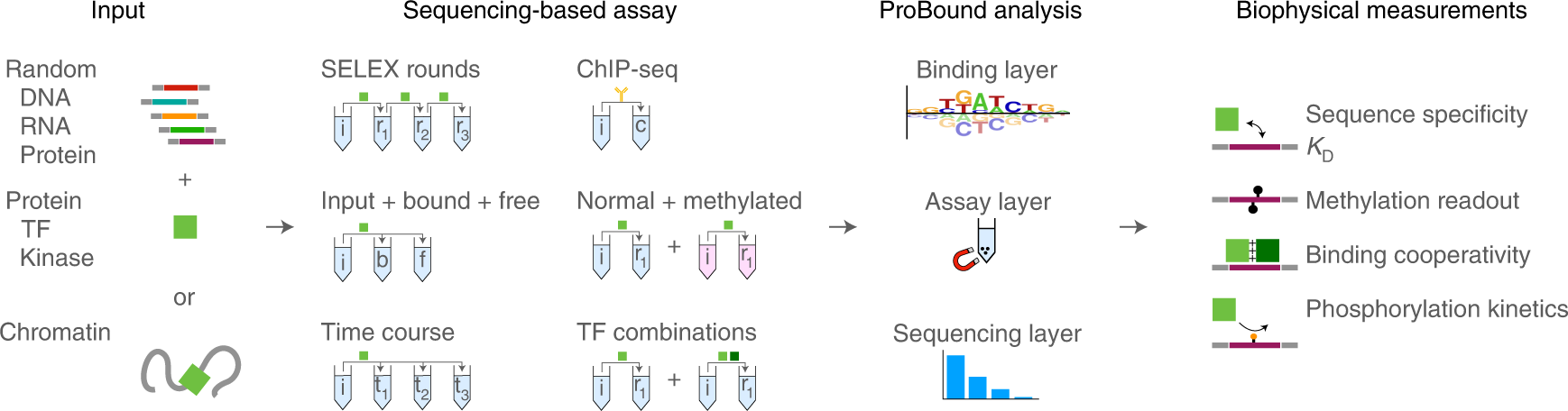
Prediction of protein–ligand binding affinity from sequencing data with interpretable machine learning | Nature Biotechnology