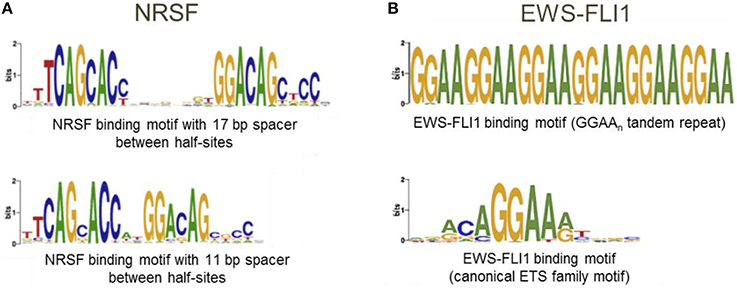
IMAGE predicts transcription factor binding sites with high confidence.... | Download Scientific Diagram

Prediction of transcription factor bindings sites affected by SNPs located at the osteopontin promoter - ScienceDirect
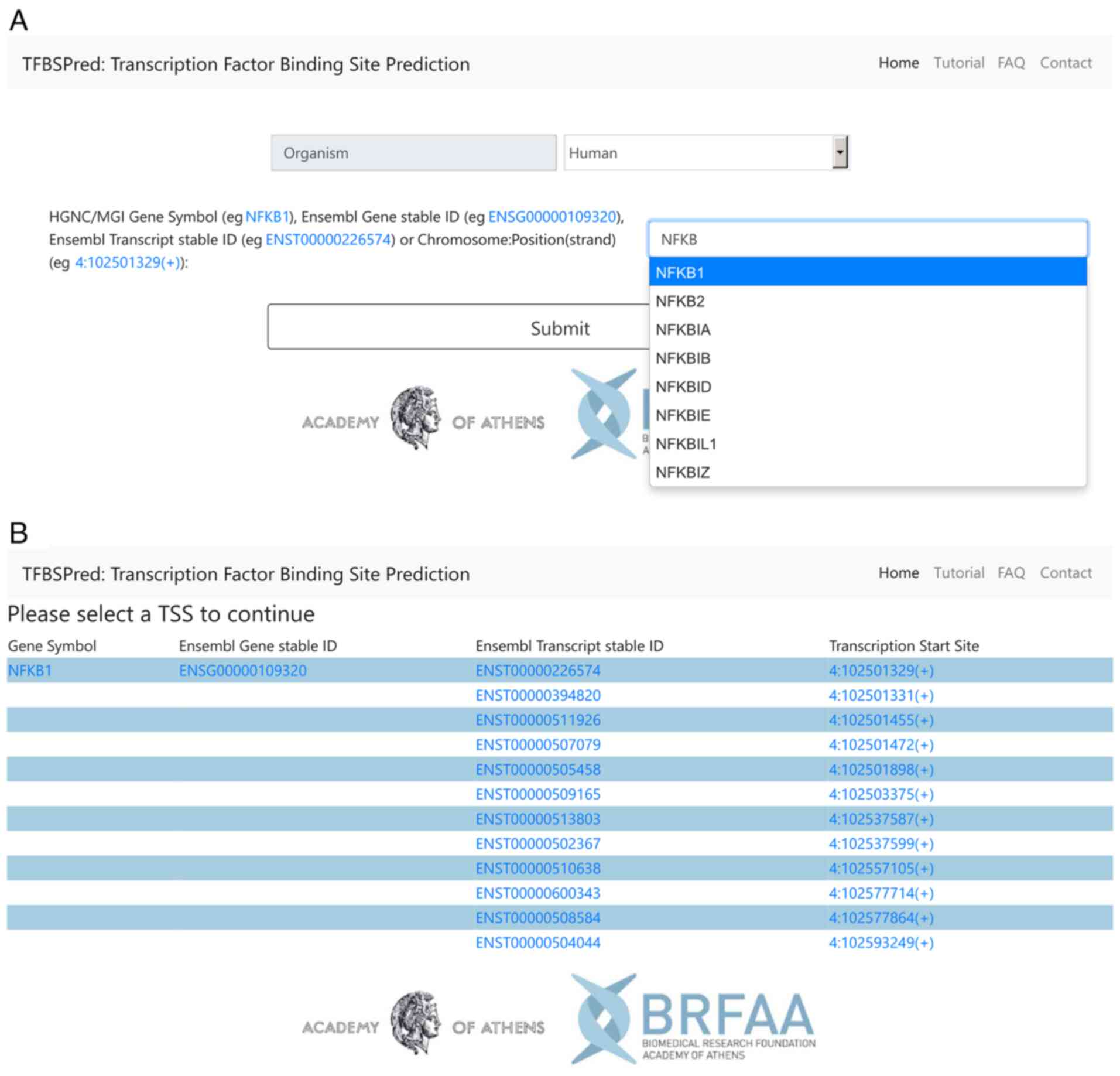
IJMS | Free Full-Text | Genetic Variants in Transcription Factor Binding Sites in Humans: Triggered by Natural Selection and Triggers of Diseases

Exploiting transcription factor binding site clustering to identify cis-regulatory modules involved in pattern formation in the Drosophila genome | PNAS
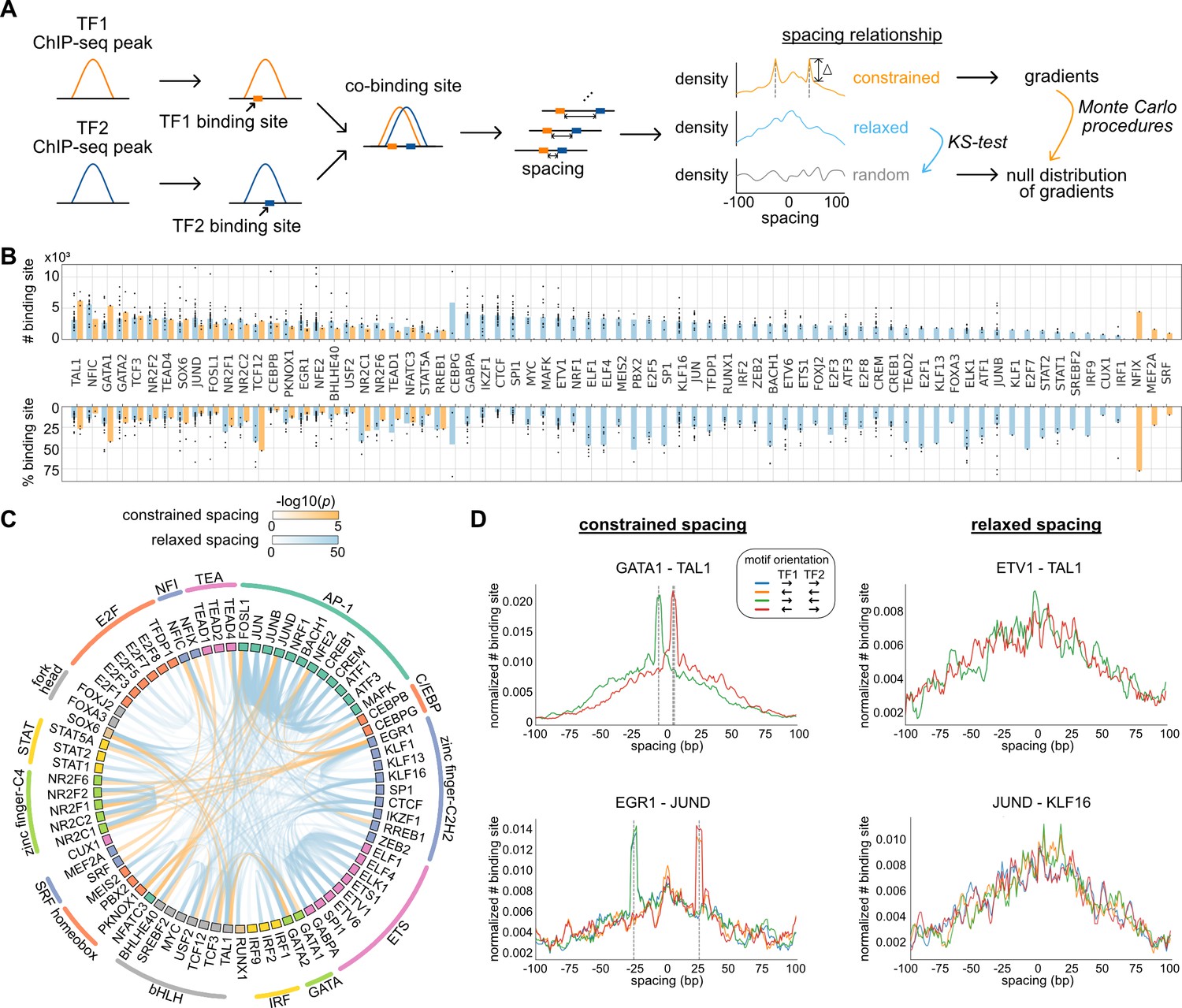
Systematic analysis of naturally occurring insertions and deletions that alter transcription factor spacing identifies tolerant and sensitive transcription factor pairs | eLife

Prediction of transcription factor binding sites and mutation analysis... | Download Scientific Diagram

Prediction of transcription factor bindings sites affected by SNPs located at the osteopontin promoter - ScienceDirect
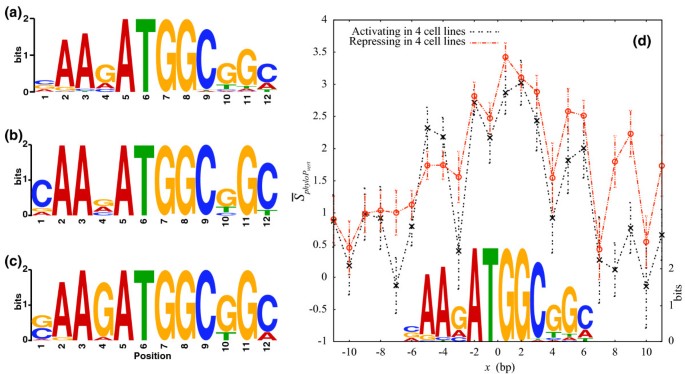
Functional analysis of transcription factor binding sites in human promoters | Genome Biology | Full Text

Discovering unknown human and mouse transcription factor binding sites and their characteristics from ChIP-seq data | PNAS
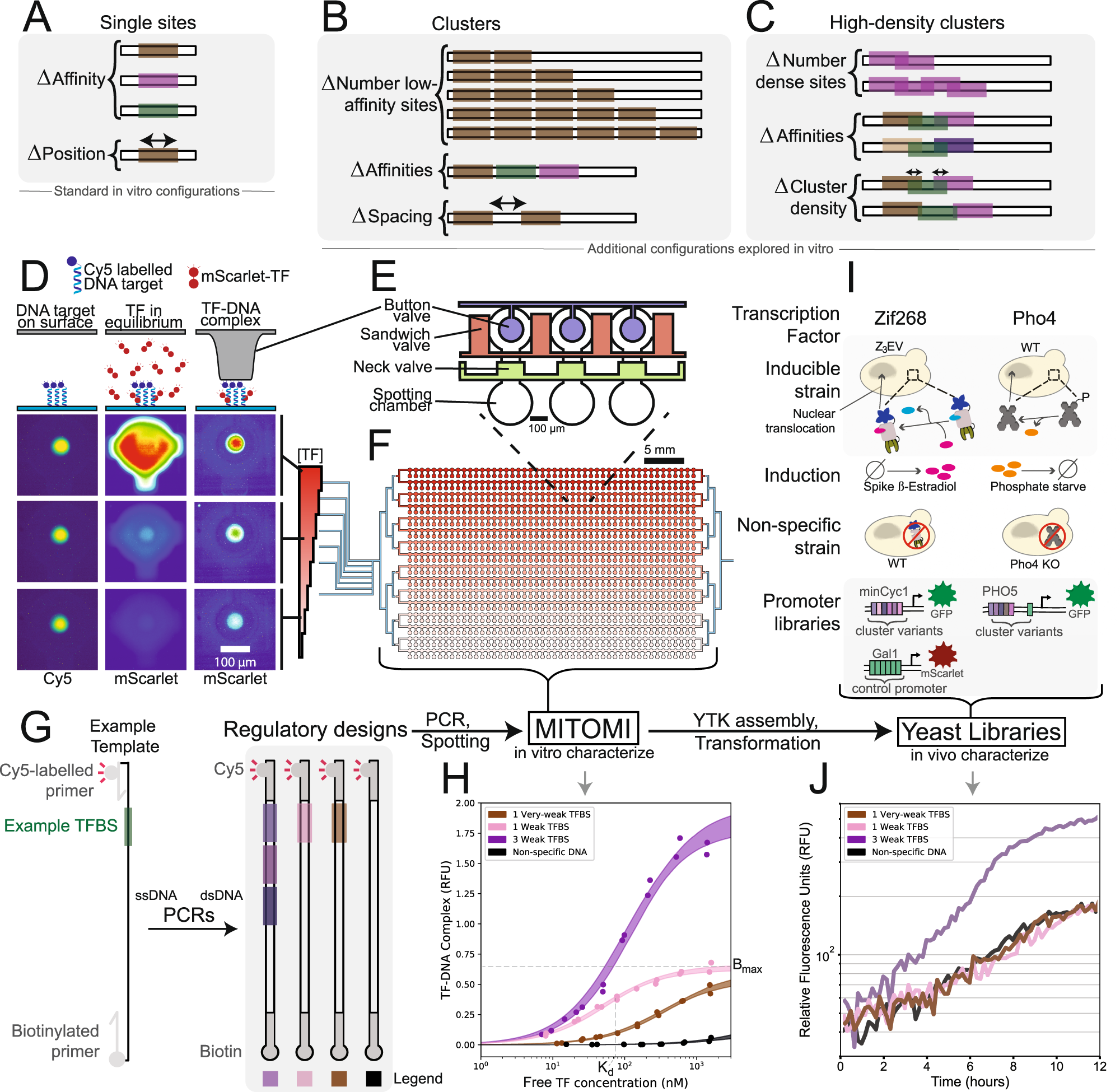
Systematic analysis of low-affinity transcription factor binding site clusters in vitro and in vivo establishes their functional relevance | Nature Communications