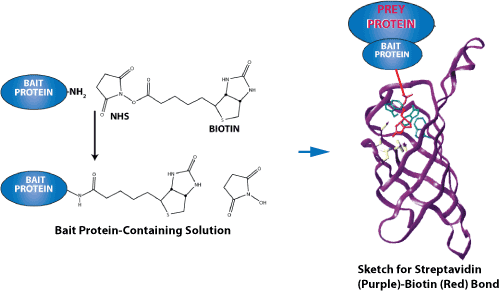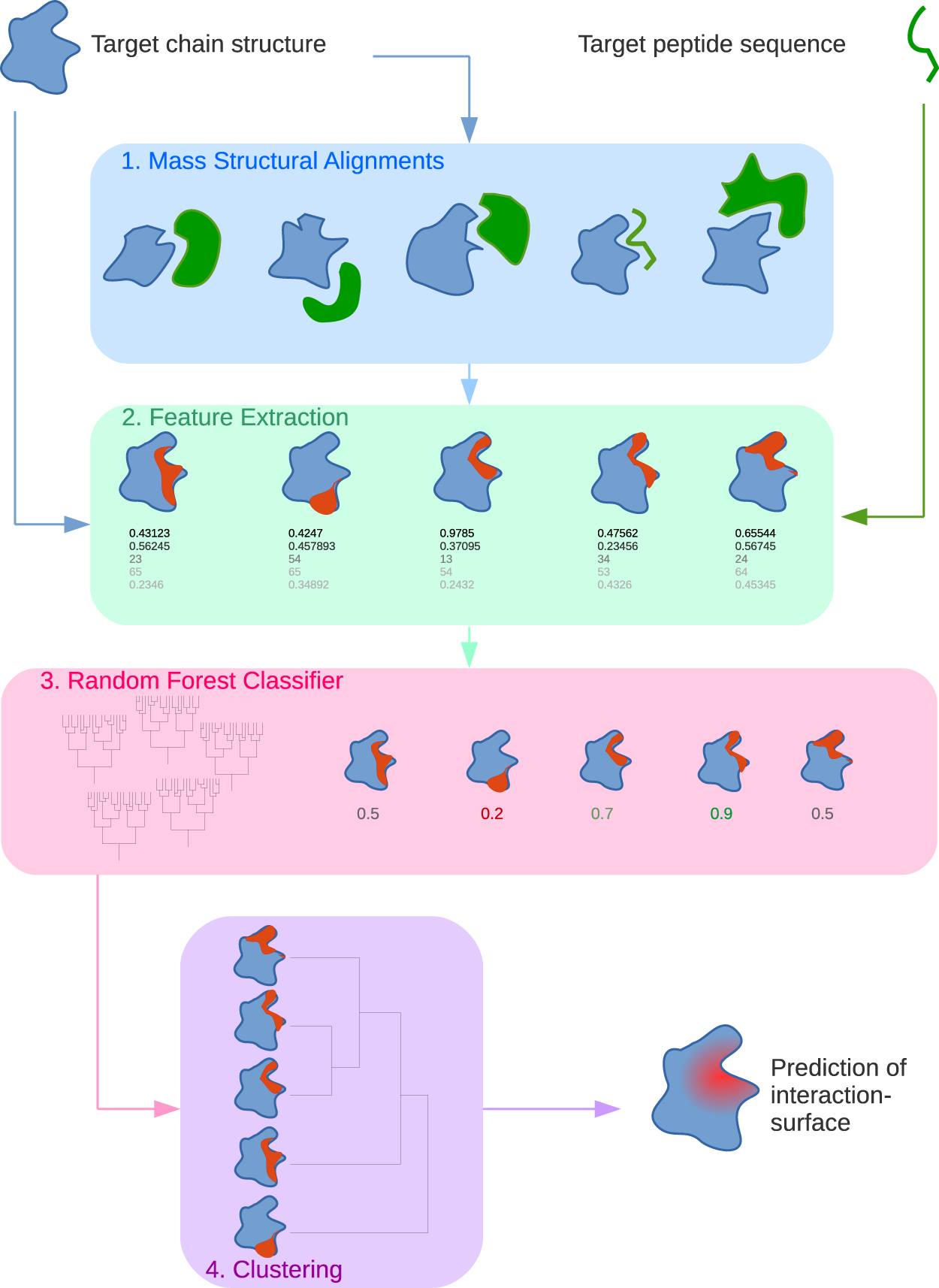
Predicting protein-peptide interaction sites using distant protein complexes as structural templates | Scientific Reports
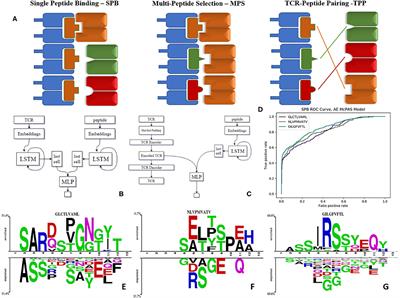
Schematics depicting different binding mechanisms for peptide–protein... | Download Scientific Diagram
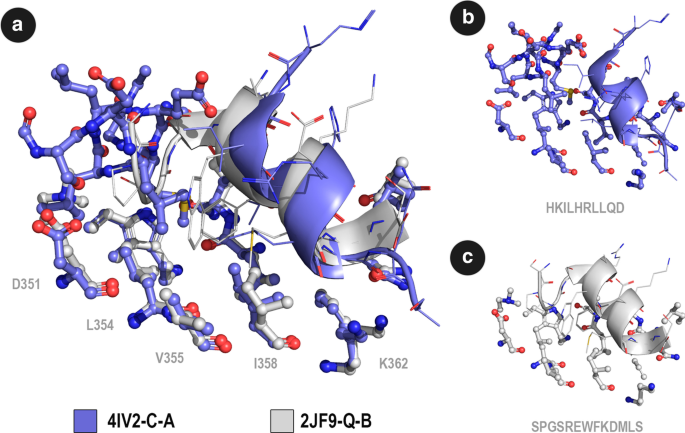
Propedia: a database for protein–peptide identification based on a hybrid clustering algorithm | BMC Bioinformatics | Full Text

Structural and computational analysis of peptide recognition mechanism of class-C type penicillin binding protein, alkaline D-peptidase from Bacillus cereus DF4-B | Scientific Reports

Protein–Peptide Binding Site Detection Using 3D Convolutional Neural Networks | Journal of Chemical Information and Modeling

Structure‐Based Design of Inhibitors of Protein–Protein Interactions: Mimicking Peptide Binding Epitopes - Pelay‐Gimeno - 2015 - Angewandte Chemie International Edition - Wiley Online Library
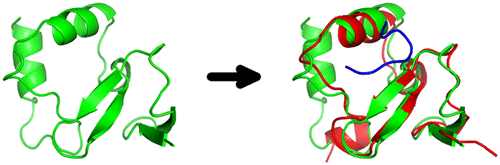
SPOT-Peptide: Template-Based Prediction of Peptide-Binding Proteins and Peptide-Binding Sites.,Journal of Chemical Information and Modeling - X-MOL
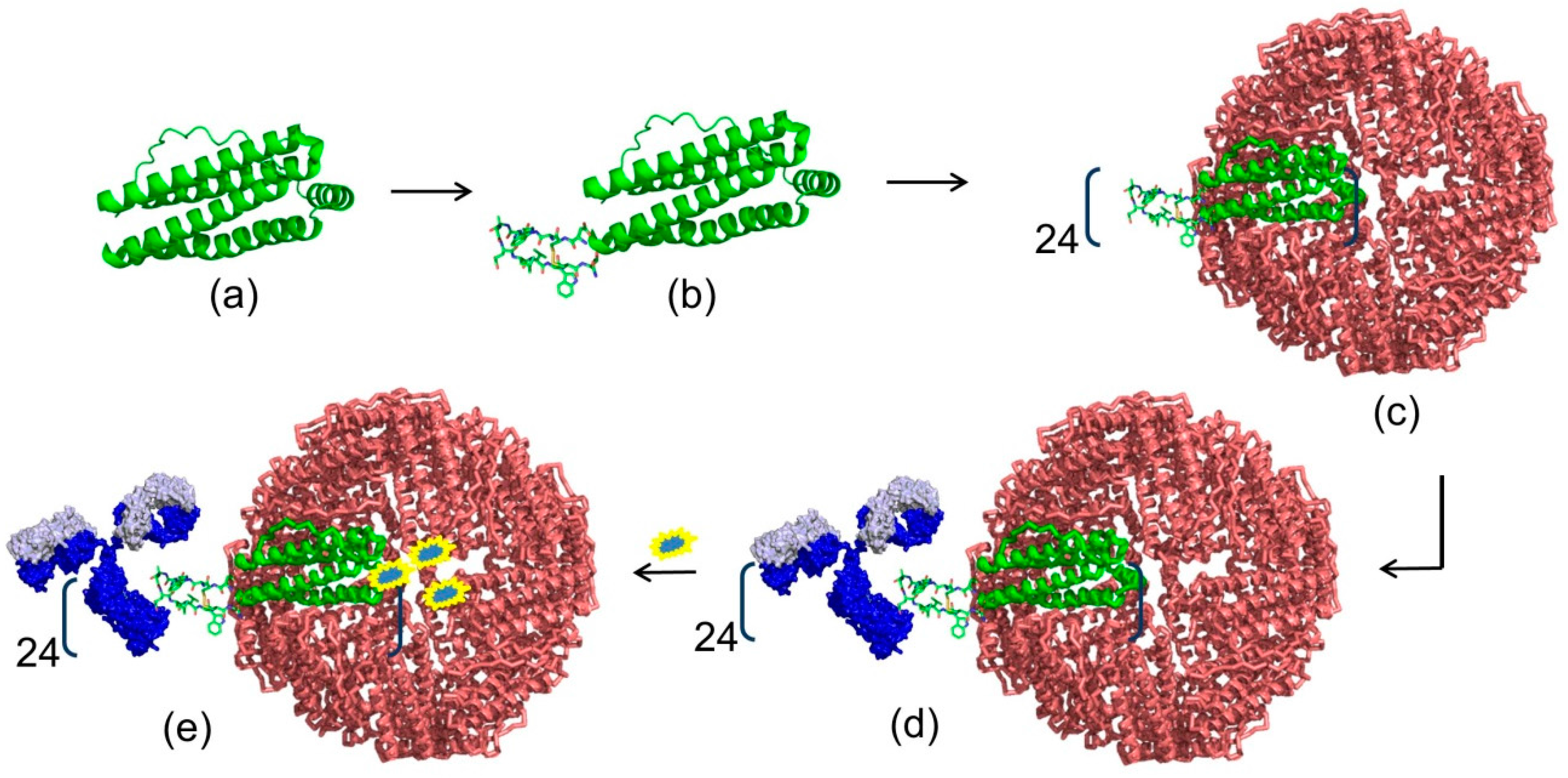
Materials | Free Full-Text | Fc-Binding Ligands of Immunoglobulin G: An Overview of High Affinity Proteins and Peptides

Investigation of Penicillin Binding Protein (PBP)-like Peptide Cyclase and Hydrolase in Surugamide Non-ribosomal Peptide Biosynthesis - ScienceDirect
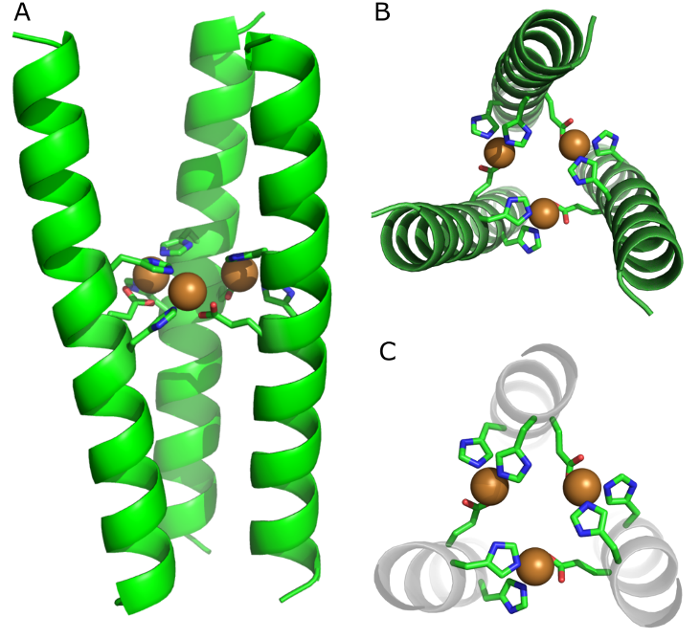
Self-assembly properties and applications of metal-binding peptides and proteins - Leiden University

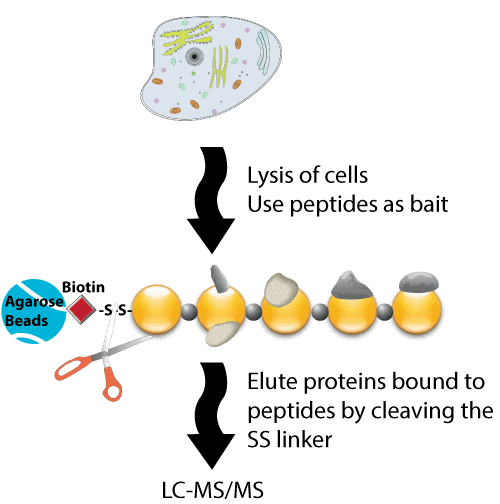
Mapping low-affinity/high-specificity peptide–protein interactions using ligand-footprinting mass spectrometry | PNAS

A Polymorphic Pocket at the P10 Position Contributes to Peptide Binding Specificity in Class II MHC Proteins: Chemistry & Biology