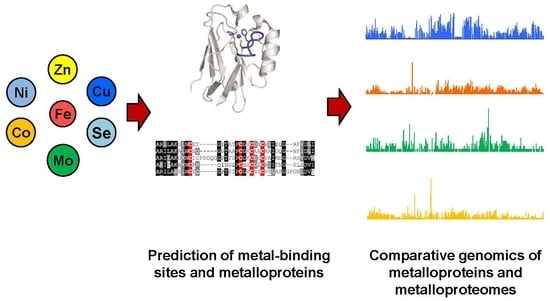
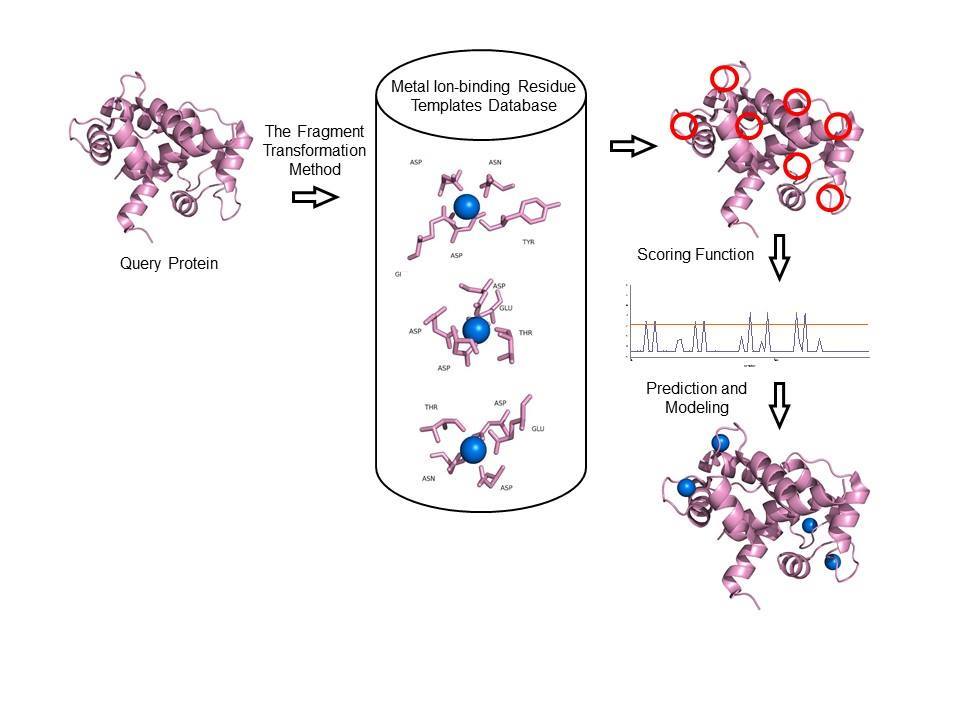
Prediction of Metal Ion–Binding Sites in Proteins Using the Fragment Transformation Method | PLOS ONE

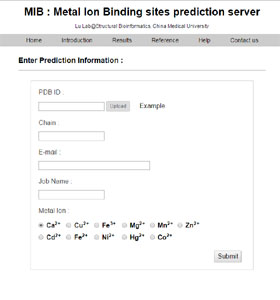
MIB: Metal Ion-Binding Site Prediction and Docking Server | Journal of Chemical Information and Modeling

Prediction of water and metal binding sites and their affinities by using the Fold-X force field | PNAS

Metal binding prediction subtasks. (a): given sequence; (b) candidate... | Download Scientific Diagram

MIB: Metal Ion-Binding Site Prediction and Docking Server | Journal of Chemical Information and Modeling
GitHub - sbl-sdsc/metal-binding-prediction: Methods to predict metal binding sites in a protein by its amino acid sequence
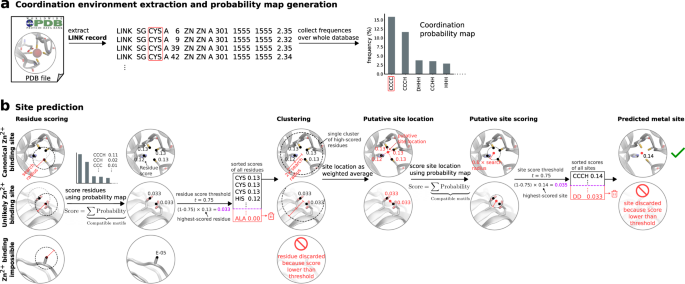
Metal3D: a general deep learning framework for accurate metal ion location prediction in proteins | Nature Communications

Alignment-free metal ion-binding site prediction from protein sequence through pretrained language model and multi-task learning | bioRxiv
Prediction of Metal Ion–Binding Sites in Proteins Using the Fragment Transformation Method | PLOS ONE
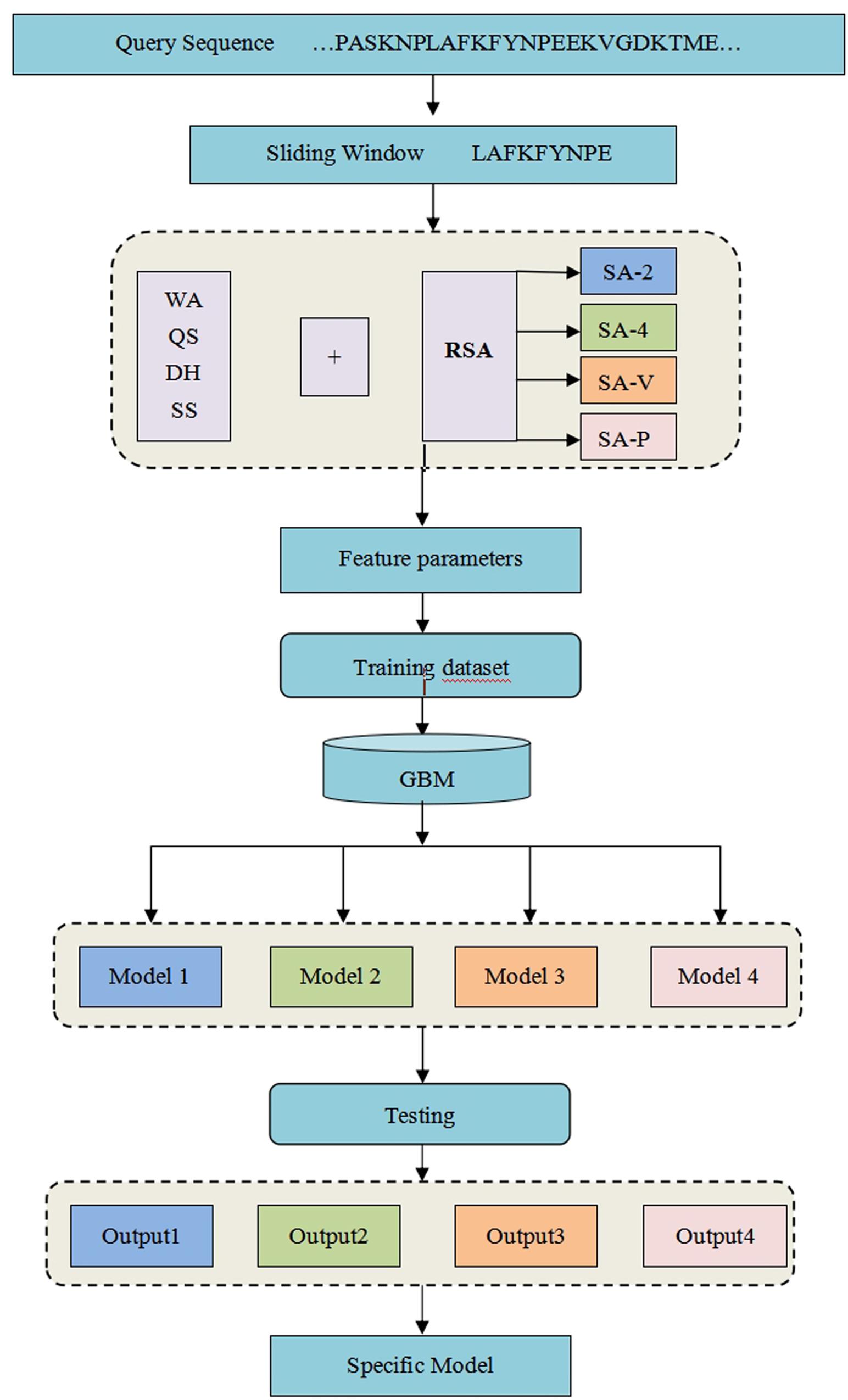
Frontiers | The Identification of Metal Ion Ligand-Binding Residues by Adding the Reclassified Relative Solvent Accessibility
GitHub - biomed-AI/LMetalSite: MetalSite: alignment-free metal ion-binding site prediction from protein sequence through pretrained language model and multi-task learning

Exploring the computational methods for protein-ligand binding site prediction - Computational and Structural Biotechnology Journal
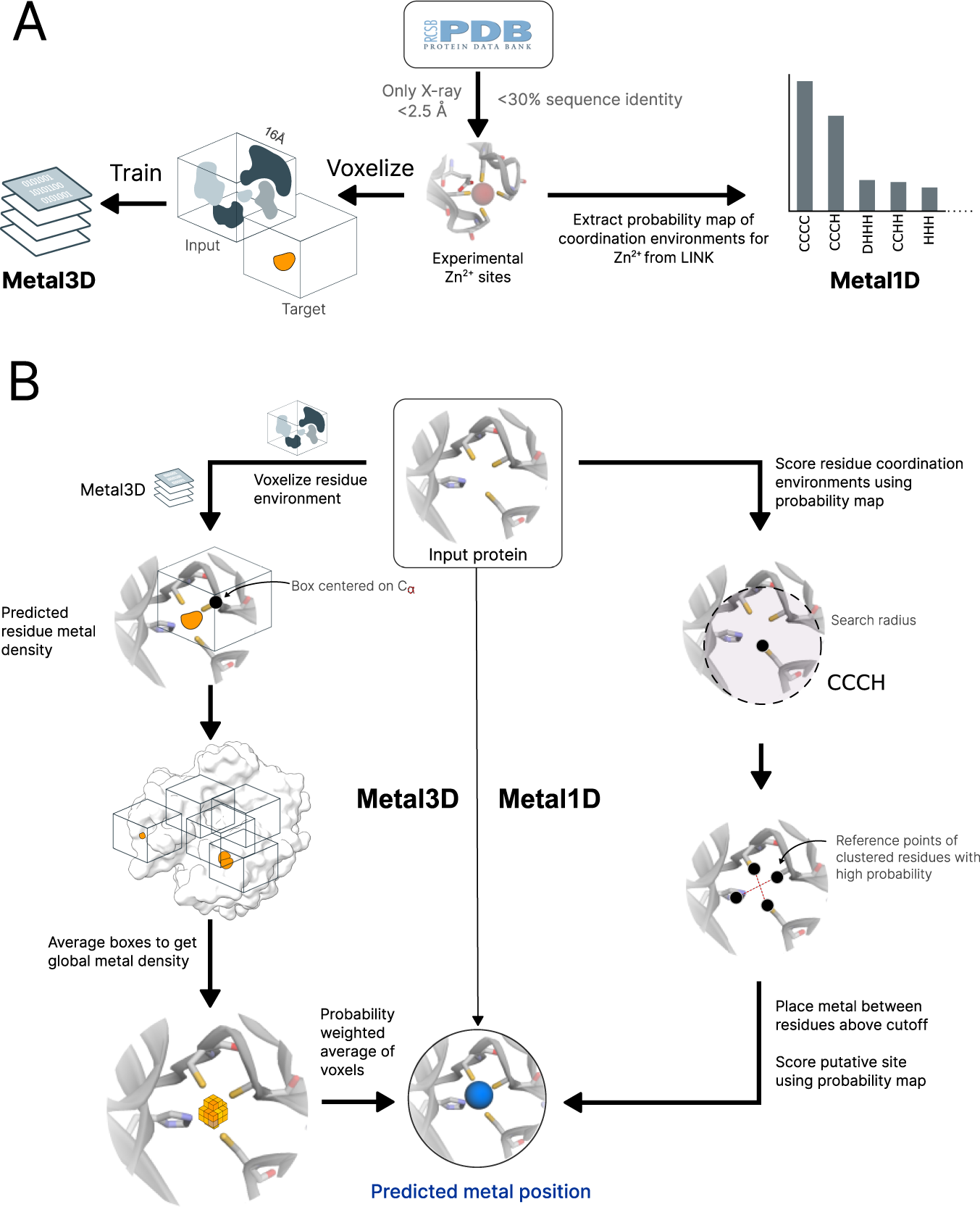
Metal3D: a general deep learning framework for accurate metal ion location prediction in proteins | Nature Communications

PRL-3 has numerous predicted metal binding sites. The crystal structure... | Download Scientific Diagram