Learning non-stationary Langevin dynamics from stochastic observations of latent trajectories | Nature Communications

Vivek Jayaram, John Thickstun · Parallel and Flexible Sampling from Autoregressive Models via Langevin Dynamics · SlidesLive
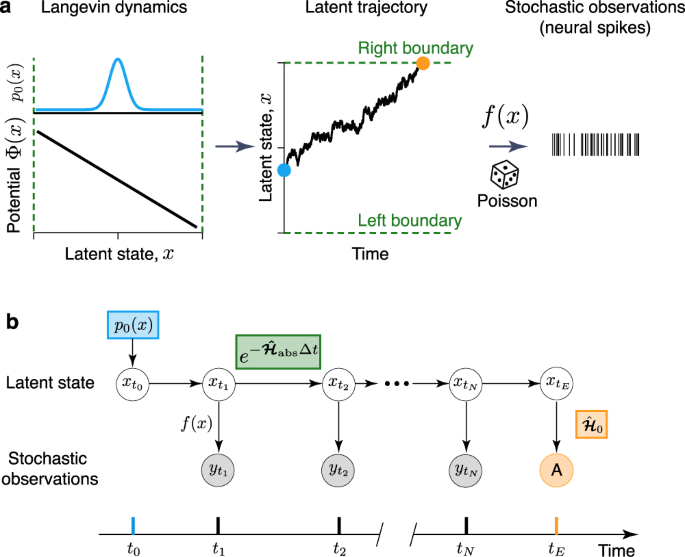
Learning non-stationary Langevin dynamics from stochastic observations of latent trajectories | Nature Communications

Generalized Langevin Equation as a Model for Barrier Crossing Dynamics in Biomolecular Folding | The Journal of Physical Chemistry B
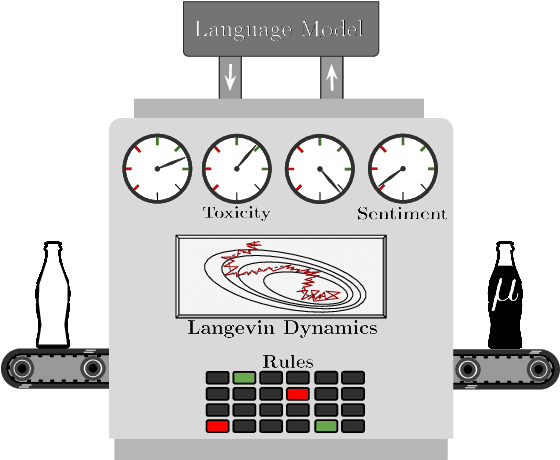
Constrained Sampling from Language Models via Langevin Dynamics in Embedding Spaces: Paper and Code - CatalyzeX

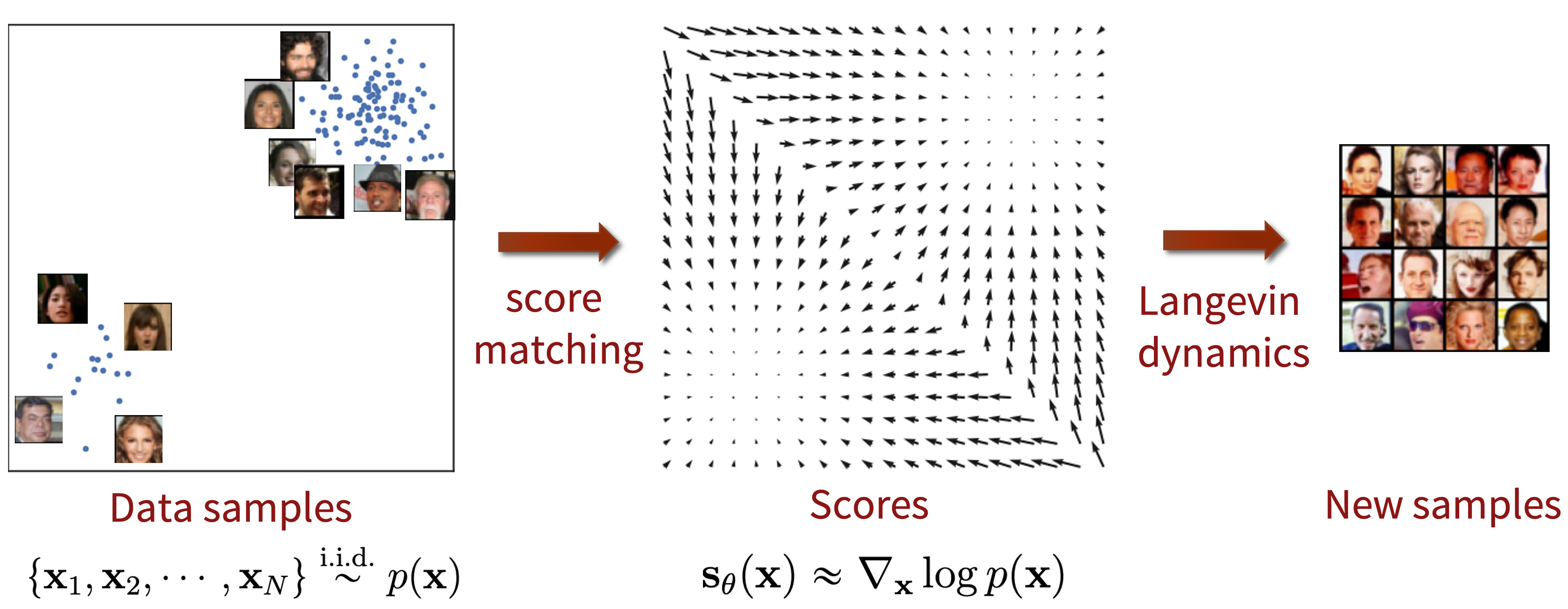
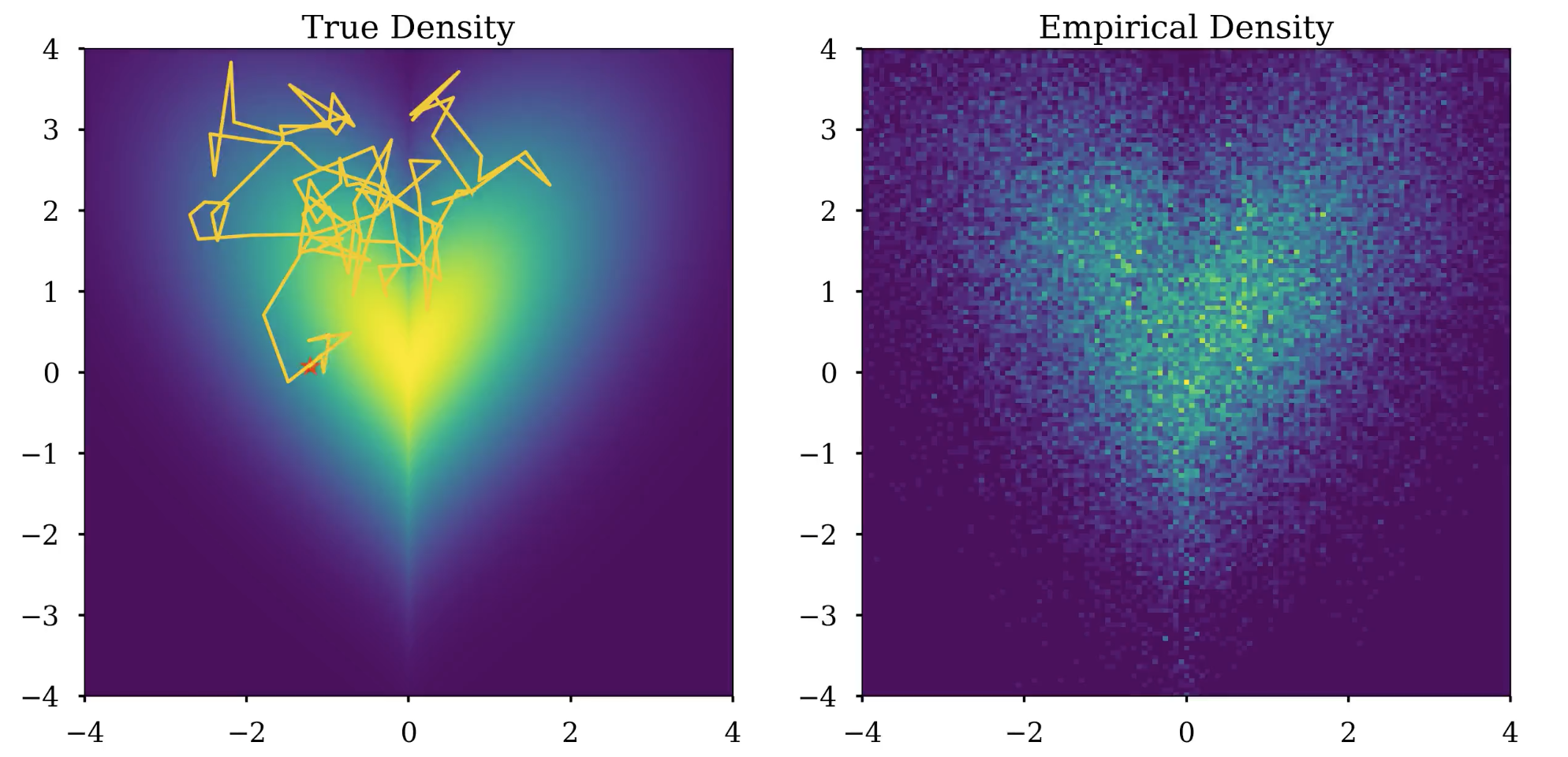
Top: Samples generated from the Langevin dynamics (7) are plotted over... | Download Scientific Diagram
![PDF] Breaking Reversibility Accelerates Langevin Dynamics for Non-Convex Optimization | Semantic Scholar PDF] Breaking Reversibility Accelerates Langevin Dynamics for Non-Convex Optimization | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/573fc80d764a90b32ef3a4a8996ed8bbc8b165d8/9-Figure1-1.png)
PDF] Breaking Reversibility Accelerates Langevin Dynamics for Non-Convex Optimization | Semantic Scholar

Understanding the Mathematical Concepts behind Langevin Dynamics | by Natan Katz | Towards Data Science

Langevin Dynamics Simulation on the Diffusivity of Polymers in Crowded Environments with Immobile Nanoparticles | Macromolecules
![PDF] Conditioned Langevin Dynamics enables efficient sampling of transition paths | Semantic Scholar PDF] Conditioned Langevin Dynamics enables efficient sampling of transition paths | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/928c7f2db6d2bd09f664cab1aa6da4cd1b0f1205/13-Figure1-1.png)
PDF] Conditioned Langevin Dynamics enables efficient sampling of transition paths | Semantic Scholar
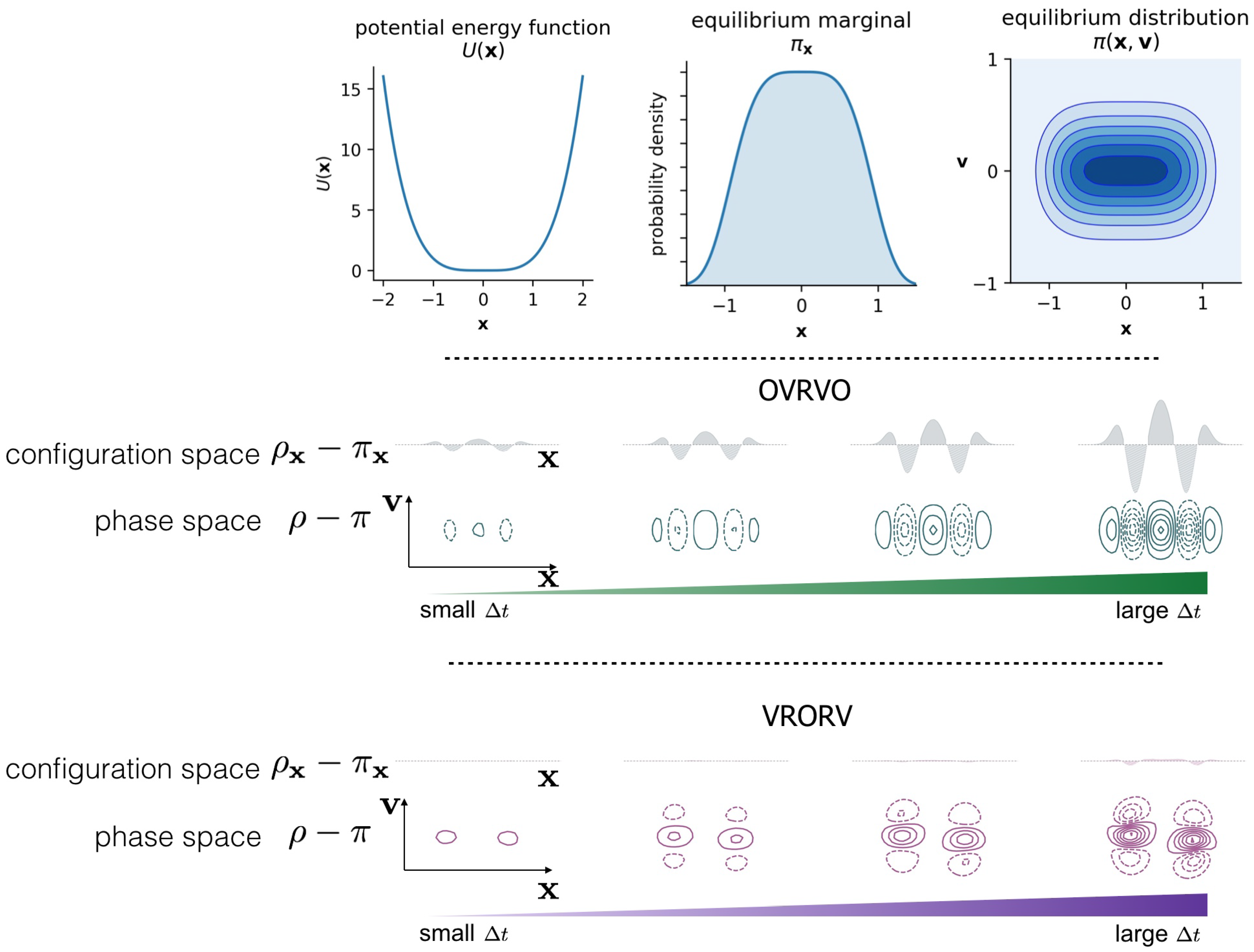
Entropy | Free Full-Text | Quantifying Configuration-Sampling Error in Langevin Simulations of Complex Molecular Systems
Illustration of Langevin dynamics. The blue lines represent different... | Download Scientific Diagram

Mathematical foundations for the Parallel Replica algorithm applied to the underdamped Langevin dynamics | MRS Communications
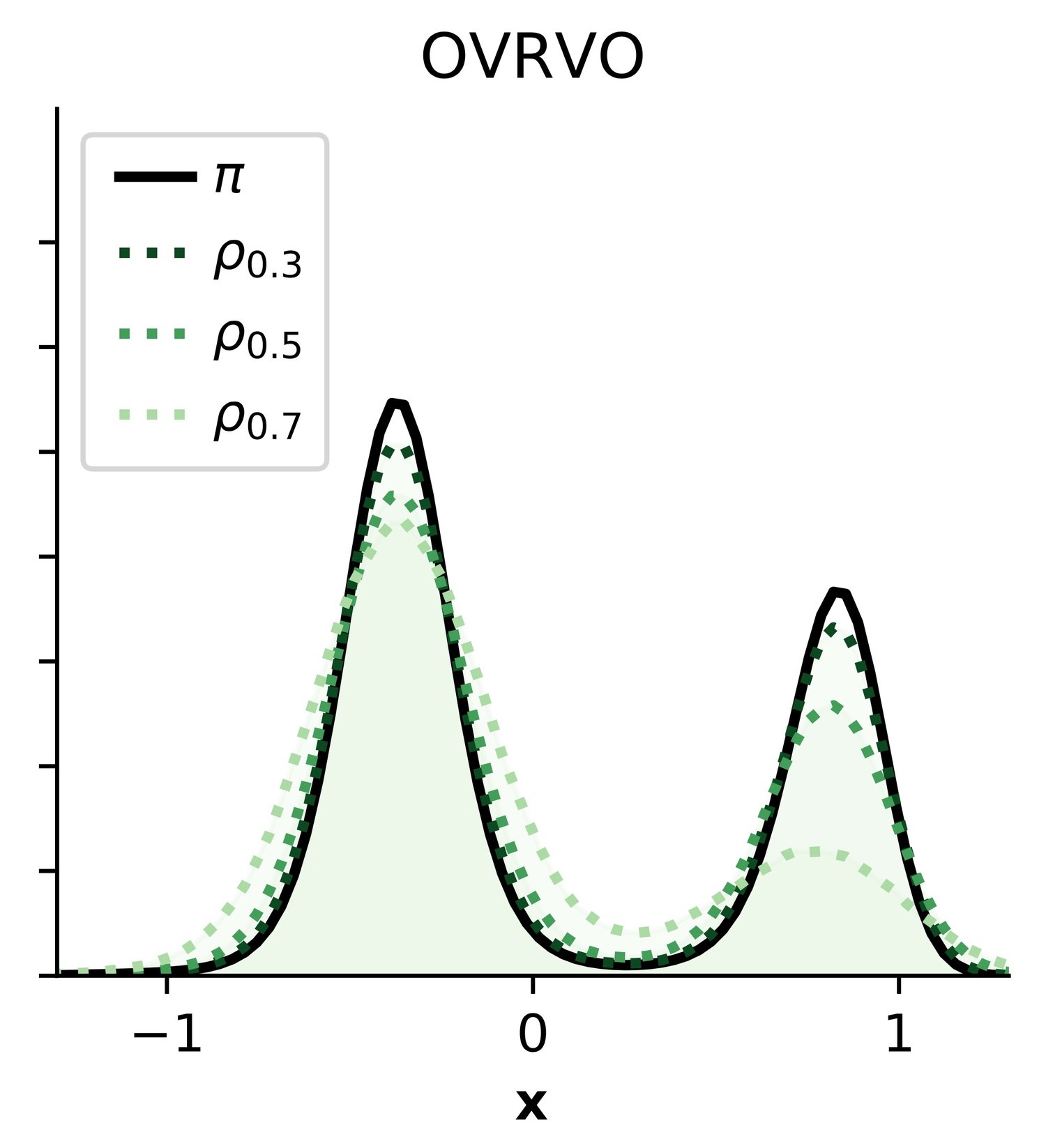
Quantifying configuration-sampling error in Langevin simulations of complex molecular systems — Chodera lab // MSKCC