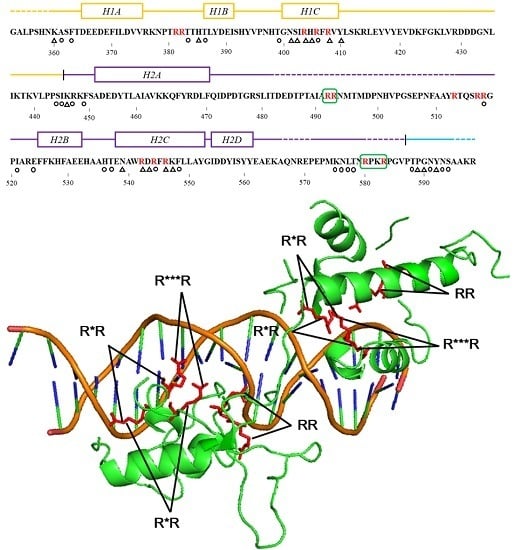
Prediction of the DNA binding and RNA-binding residues generated by... | Download Scientific Diagram

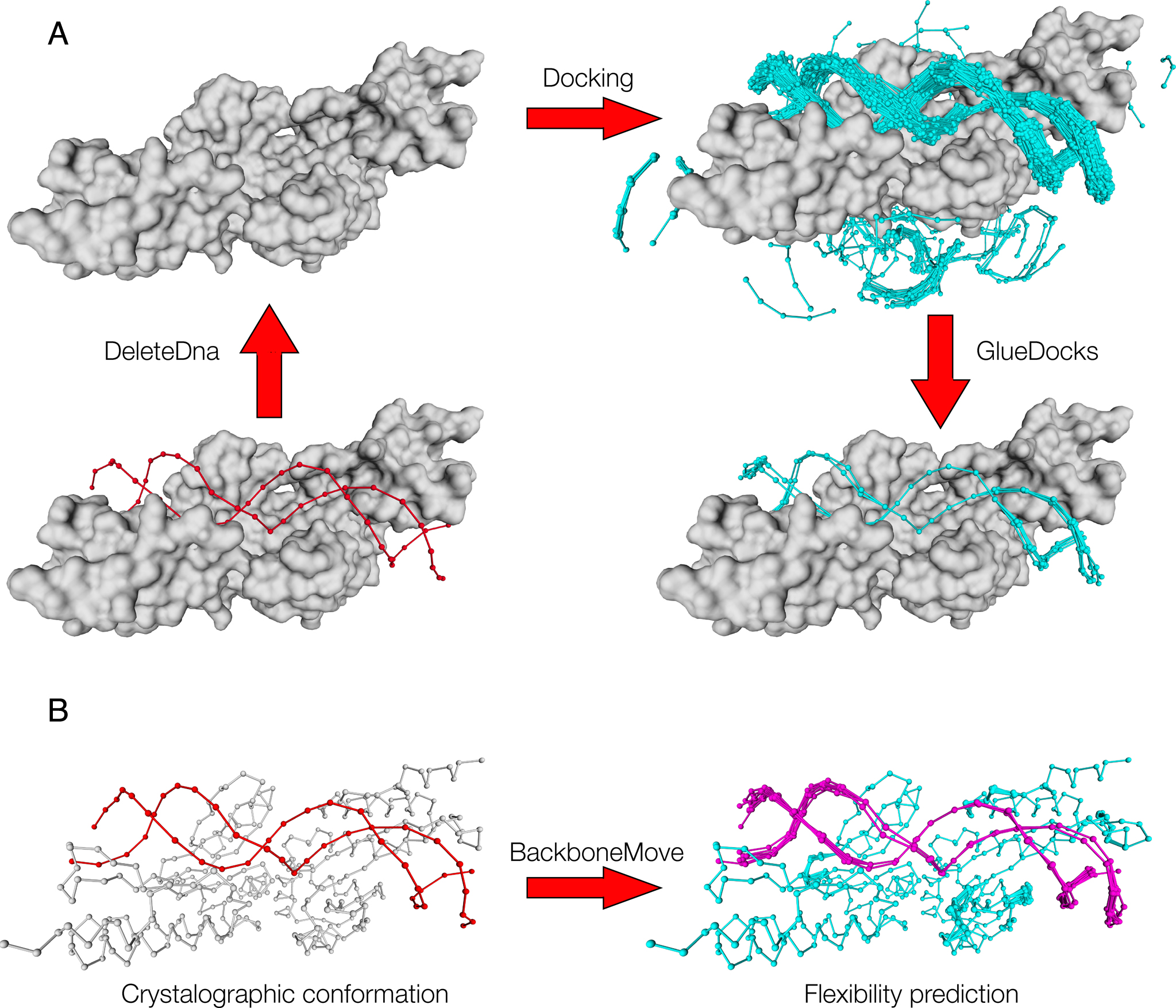
Prediction of DNA-protein binding interactions with TASSER models. (A)... | Download Scientific Diagram

Improving the prediction of DNA-protein binding by integrating multi-scale dense convolutional network with fault-tolerant coding - ScienceDirect

Predicting the DNA Binding Site Motifs of Novel Transcription Factors... | Download Scientific Diagram

Predicting transcription factor binding sites using DNA shape features based on shared hybrid deep learning architecture: Molecular Therapy - Nucleic Acids
IJMS | Free Full-Text | PSFM-DBT: Identifying DNA-Binding Proteins by Combing Position Specific Frequency Matrix and Distance-Bigram Transformation
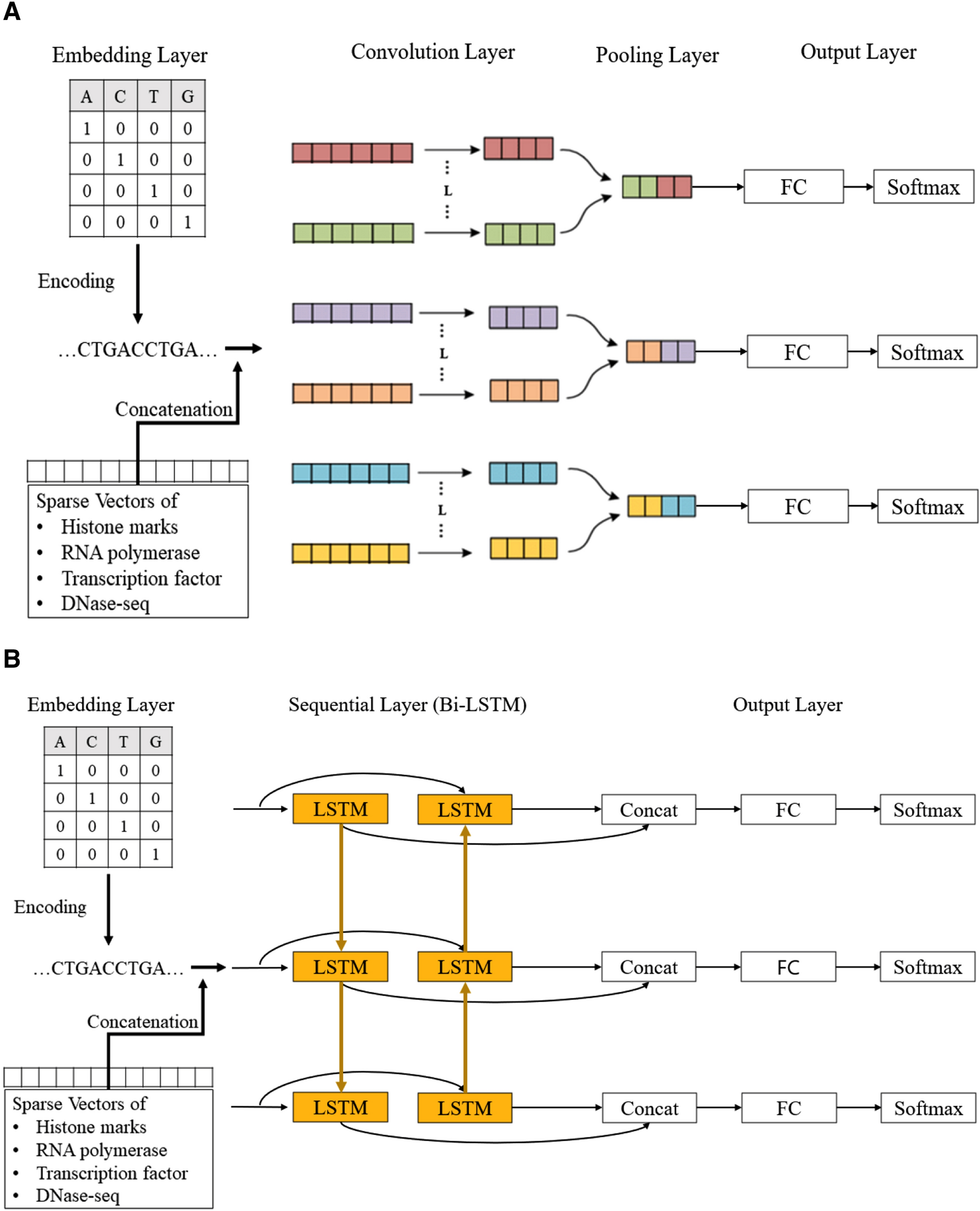
Deep learning approach for predicting functional Z-DNA regions using omics data | Scientific Reports

PDRLGB: precise DNA-binding residue prediction using a light gradient boosting machine | BMC Bioinformatics | Full Text

Overview of TF binding site prediction task. a) Transcription factors... | Download Scientific Diagram

Prediction of transcription factor binding sites and mutation analysis... | Download Scientific Diagram
![Prediction of DNA binding proteins using local features and long-term dependencies with primary sequences based on deep learning [PeerJ] Prediction of DNA binding proteins using local features and long-term dependencies with primary sequences based on deep learning [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/11262/1/fig-10-full.png)
Prediction of DNA binding proteins using local features and long-term dependencies with primary sequences based on deep learning [PeerJ]

Fasim-LongTarget enables fast and accurate genome-wide lncRNA/DNA binding prediction - Computational and Structural Biotechnology Journal

Examples of DNA-binding protein prediction on APO104. (A–D) In each... | Download Scientific Diagram
![PDF] DNA Shape Features Improve Transcription Factor Binding Site Predictions In Vivo. | Semantic Scholar PDF] DNA Shape Features Improve Transcription Factor Binding Site Predictions In Vivo. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/5d68bdf0391af25147a48171a54fea49d64d16d4/2-Figure1-1.png)
PDF] DNA Shape Features Improve Transcription Factor Binding Site Predictions In Vivo. | Semantic Scholar

Predicting Hot Spot Residues at Protein–DNA Binding Interfaces Based on Sequence Information | Interdisciplinary Sciences: Computational Life Sciences
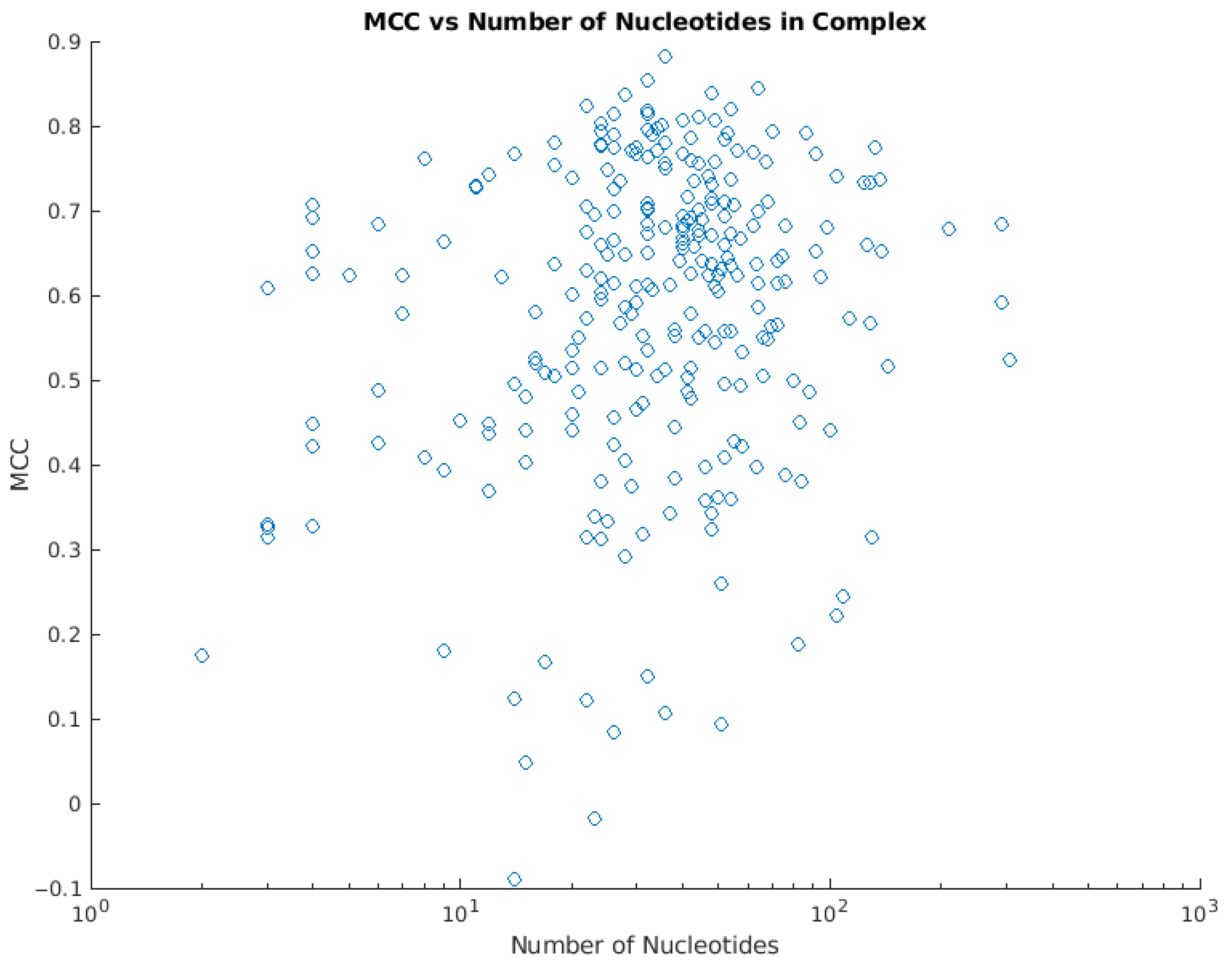
Predicting DNA-Binding Proteins and Binding Residues by Complex Structure Prediction and Application to Human Proteome | PLOS ONE
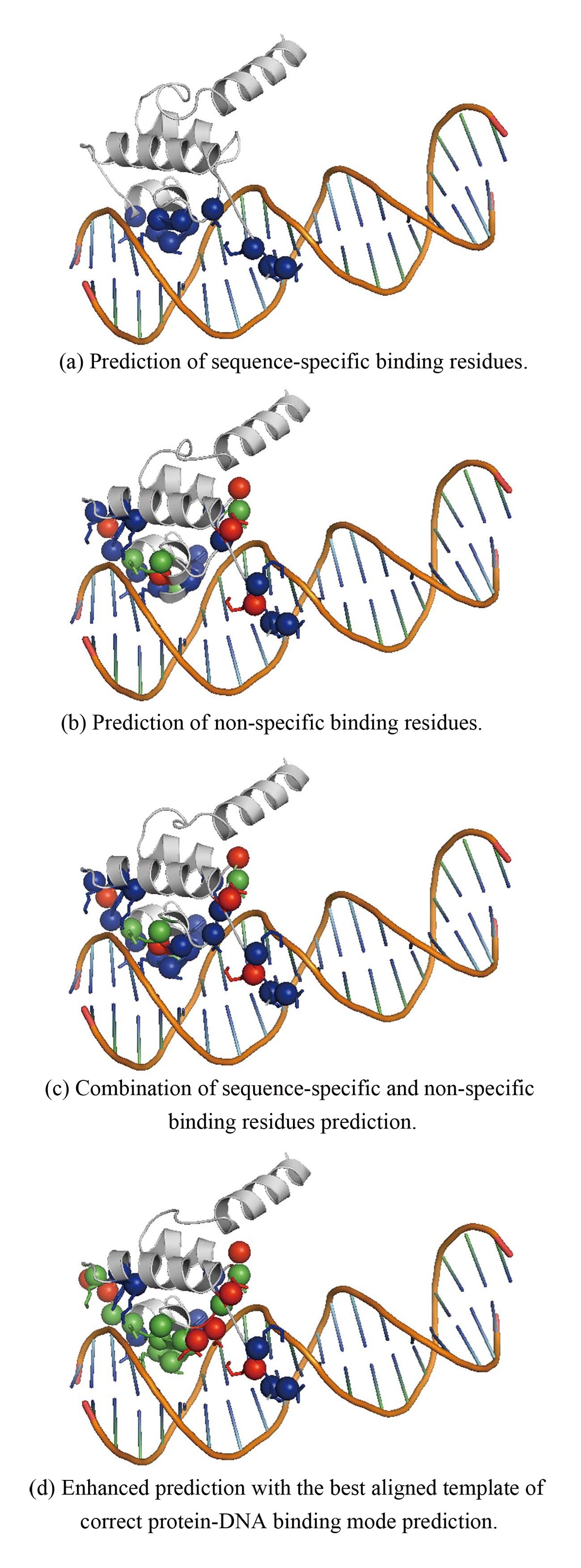
DNA-binding residues and binding mode prediction with binding-mechanism concerned models | BMC Genomics | Full Text

Prediction of DNA binding motifs from 3D models of transcription factors; identifying TLX3 regulated genes" published in Nucleic Acids Research — The Tapinos Lab