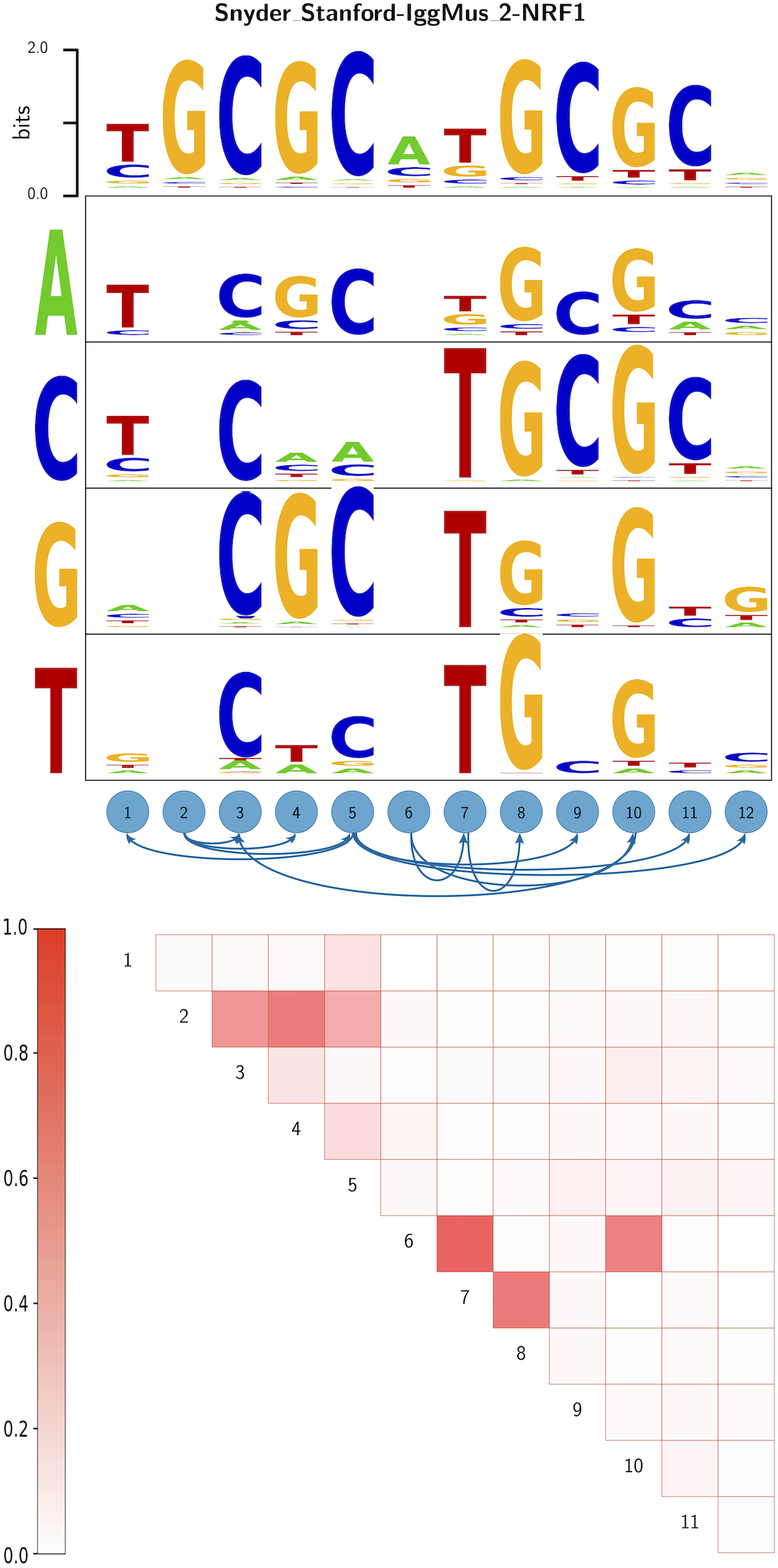
Modeling binding specificities of transcription factor pairs with random forests | BMC Bioinformatics | Full Text

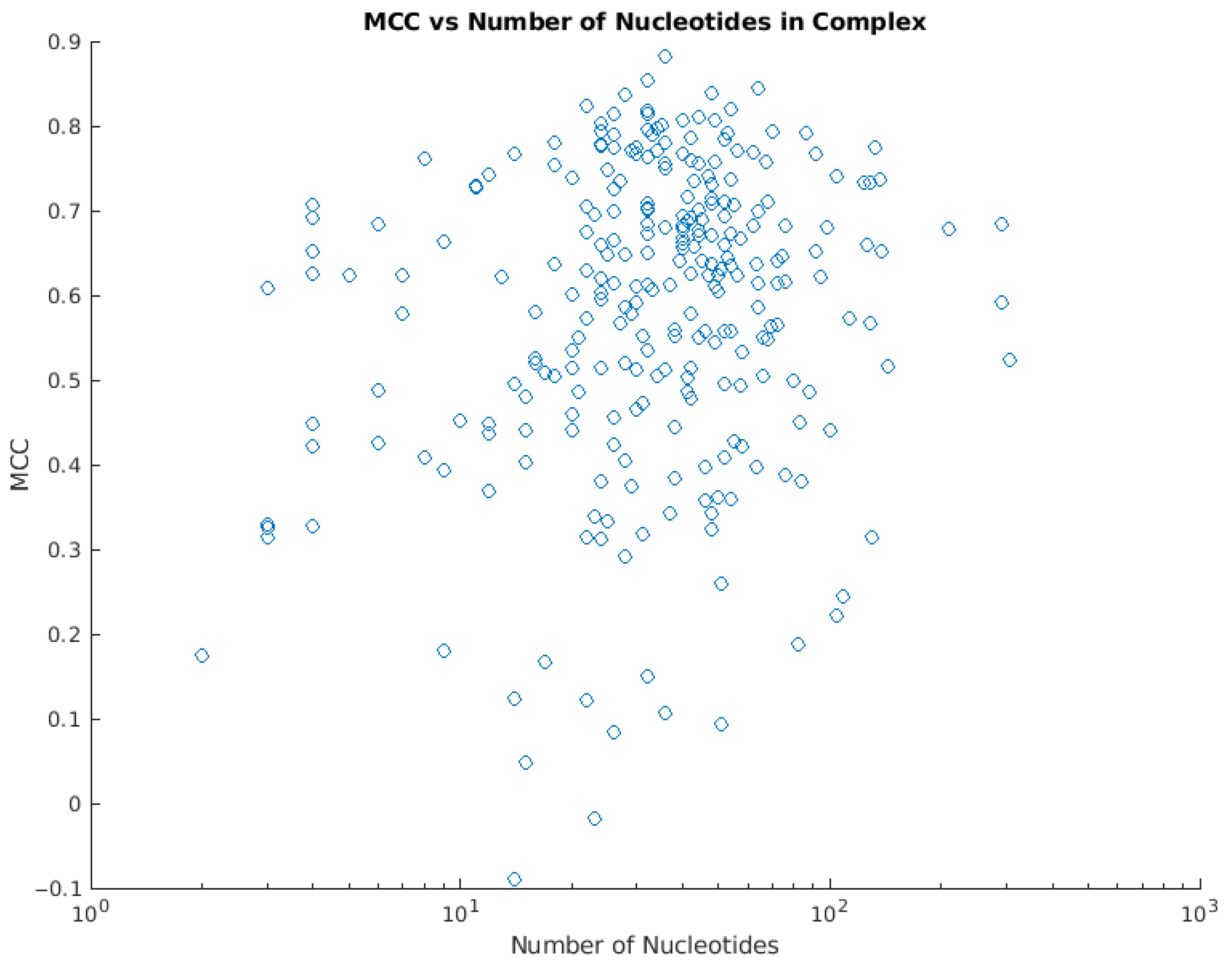
An efficient algorithm for improving structure-based prediction of transcription factor binding sites | BMC Bioinformatics | Full Text
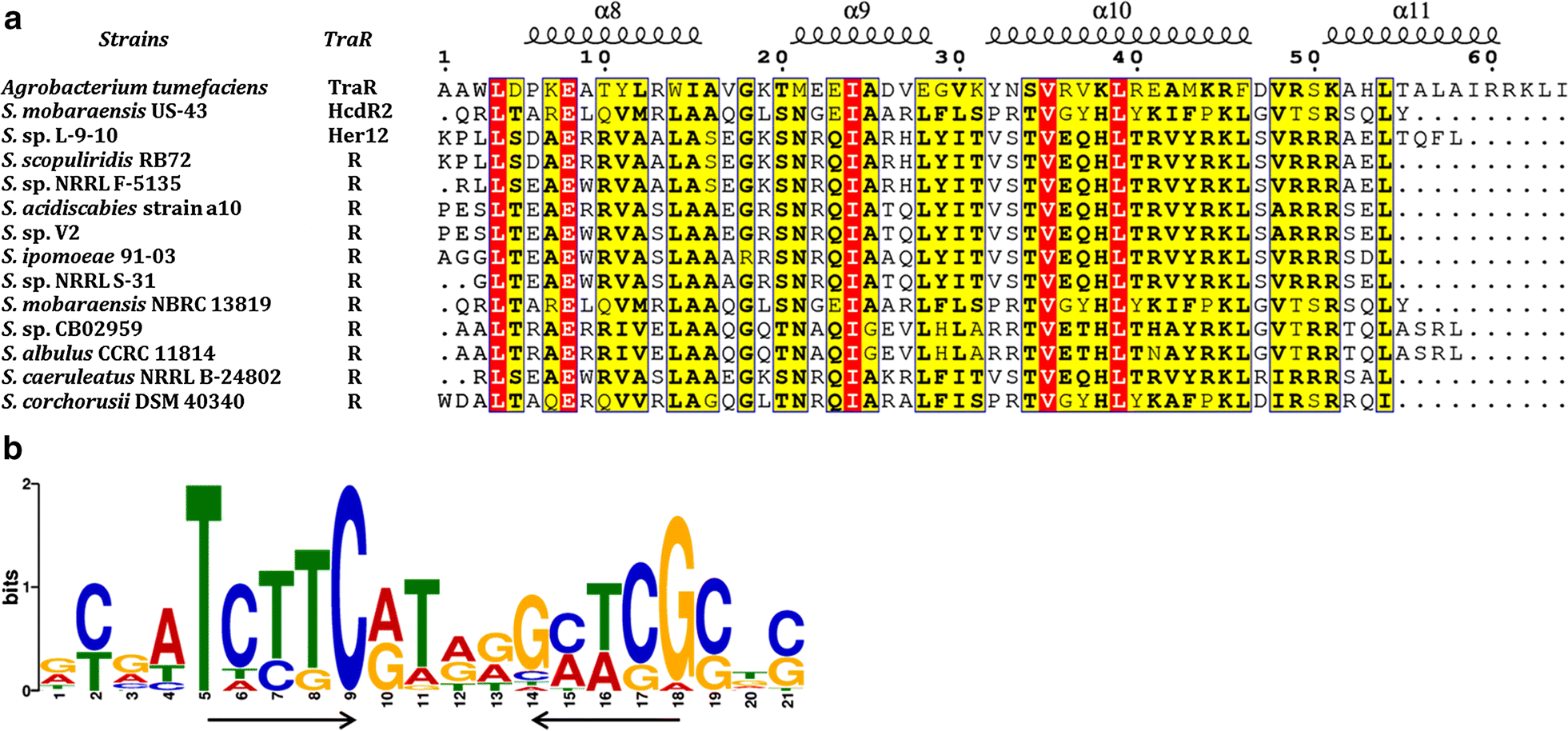

Exploring novel herbicidin analogues by transcriptional regulator overexpression and MS/MS molecular networking | Microbial Cell Factories | Full Text

Predicting transcription factor binding sites using DNA shape features based on shared hybrid deep learning architecture - ScienceDirect
GitHub - biomed-AI/GraphSite: GraphSite: protein-DNA binding site prediction using graph transformer and predicted protein structures
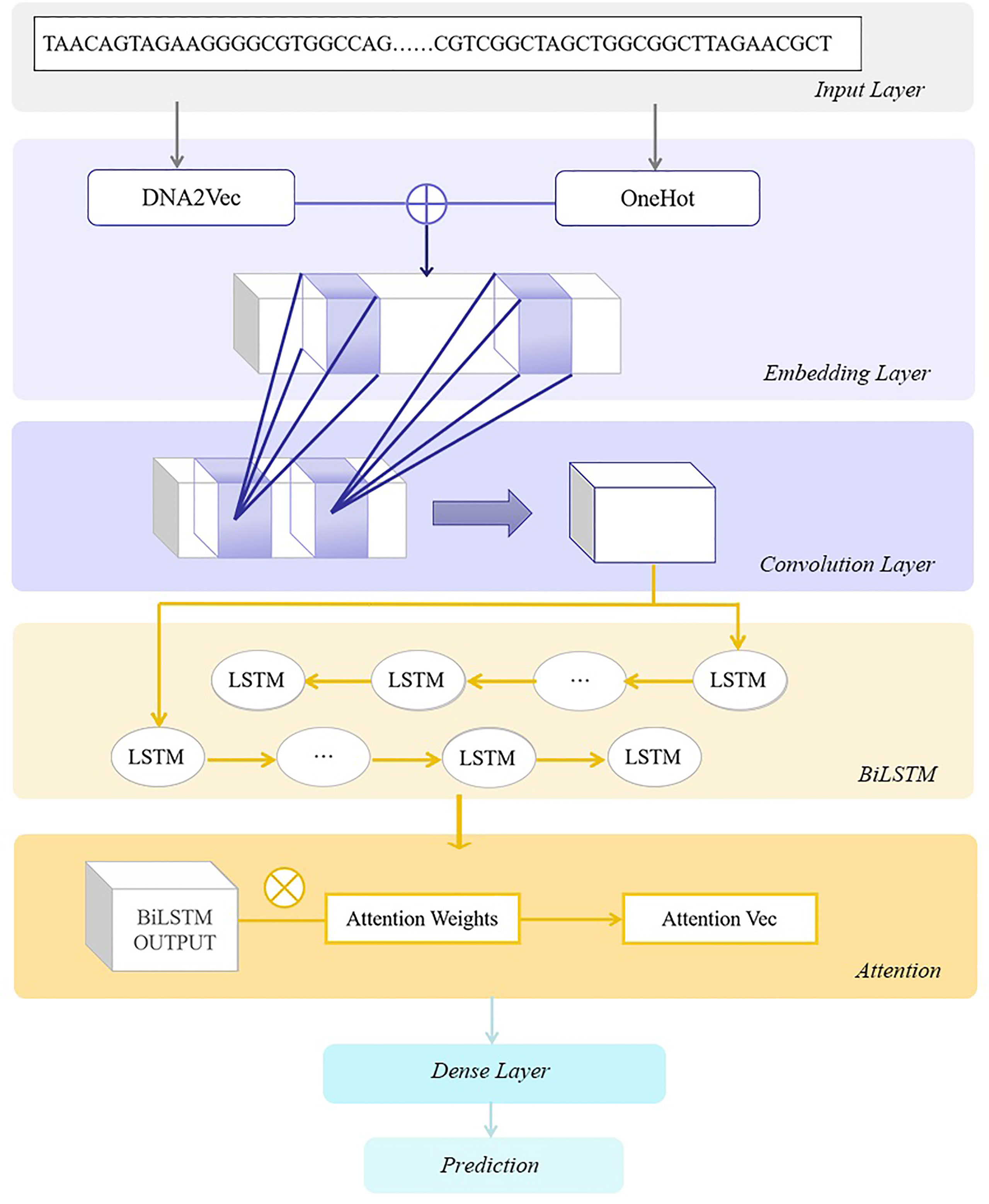
Frontiers | Prediction of Transcription Factor Binding Sites Using a Combined Deep Learning Approach
![Prediction of DNA binding proteins using local features and long-term dependencies with primary sequences based on deep learning [PeerJ] Prediction of DNA binding proteins using local features and long-term dependencies with primary sequences based on deep learning [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/11262/1/fig-1-2x.jpg)
Prediction of DNA binding proteins using local features and long-term dependencies with primary sequences based on deep learning [PeerJ]

Predicting the DNA Binding Site Motifs of Novel Transcription Factors... | Download Scientific Diagram

Prediction of transcription factor binding sites and mutation analysis... | Download Scientific Diagram

Examples of DNA-binding protein prediction on APO104. (A–D) In each... | Download Scientific Diagram
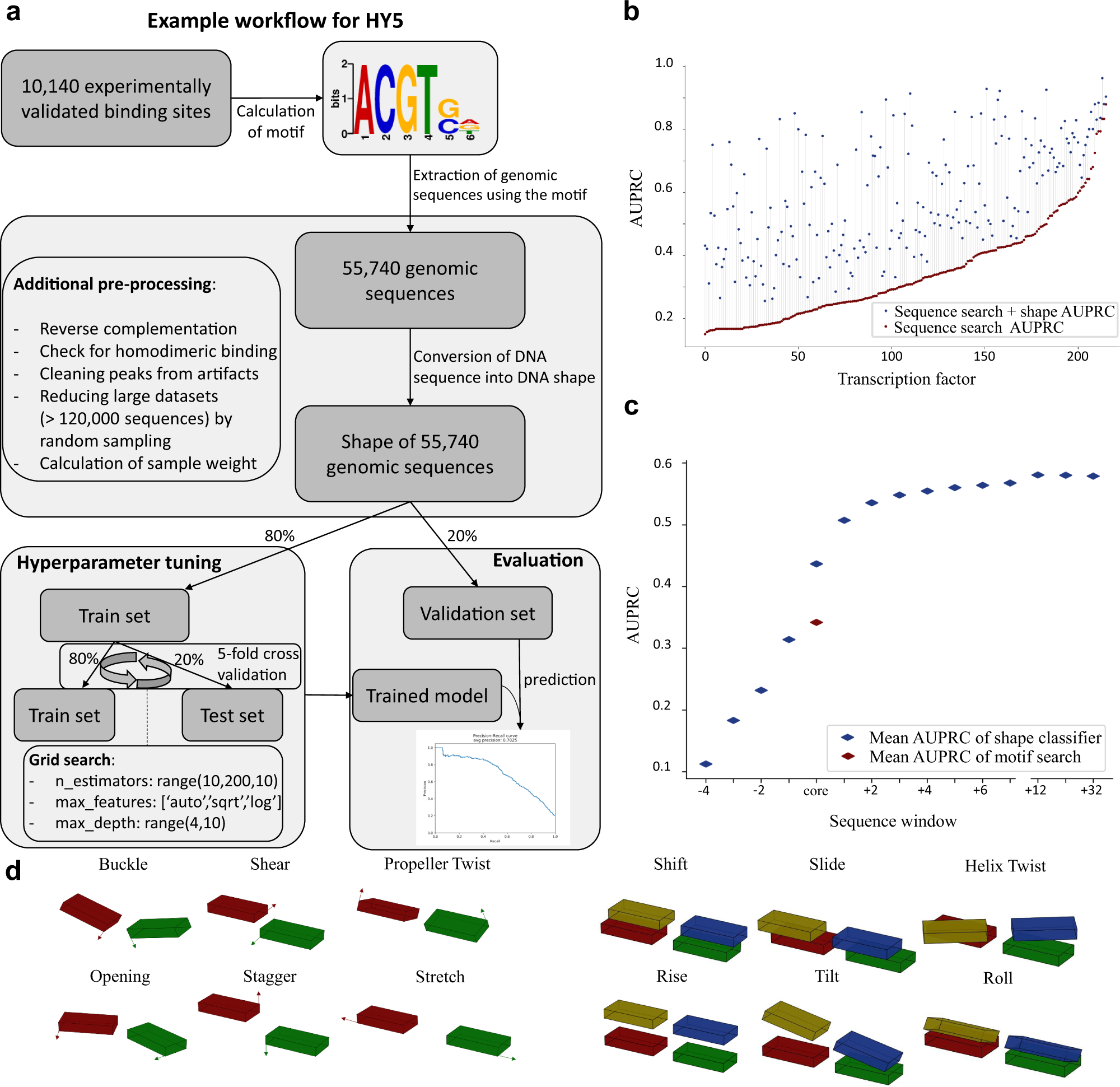
Local DNA shape is a general principle of transcription factor binding specificity in Arabidopsis thaliana | Nature Communications

Improving DNA-Binding Protein Prediction Using Three-Part Sequence-Order Feature Extraction and a Deep Neural Network Algorithm | Journal of Chemical Information and Modeling

Prediction results for putative NLS, NES and DNA-binding motifs on CAV... | Download Scientific Diagram
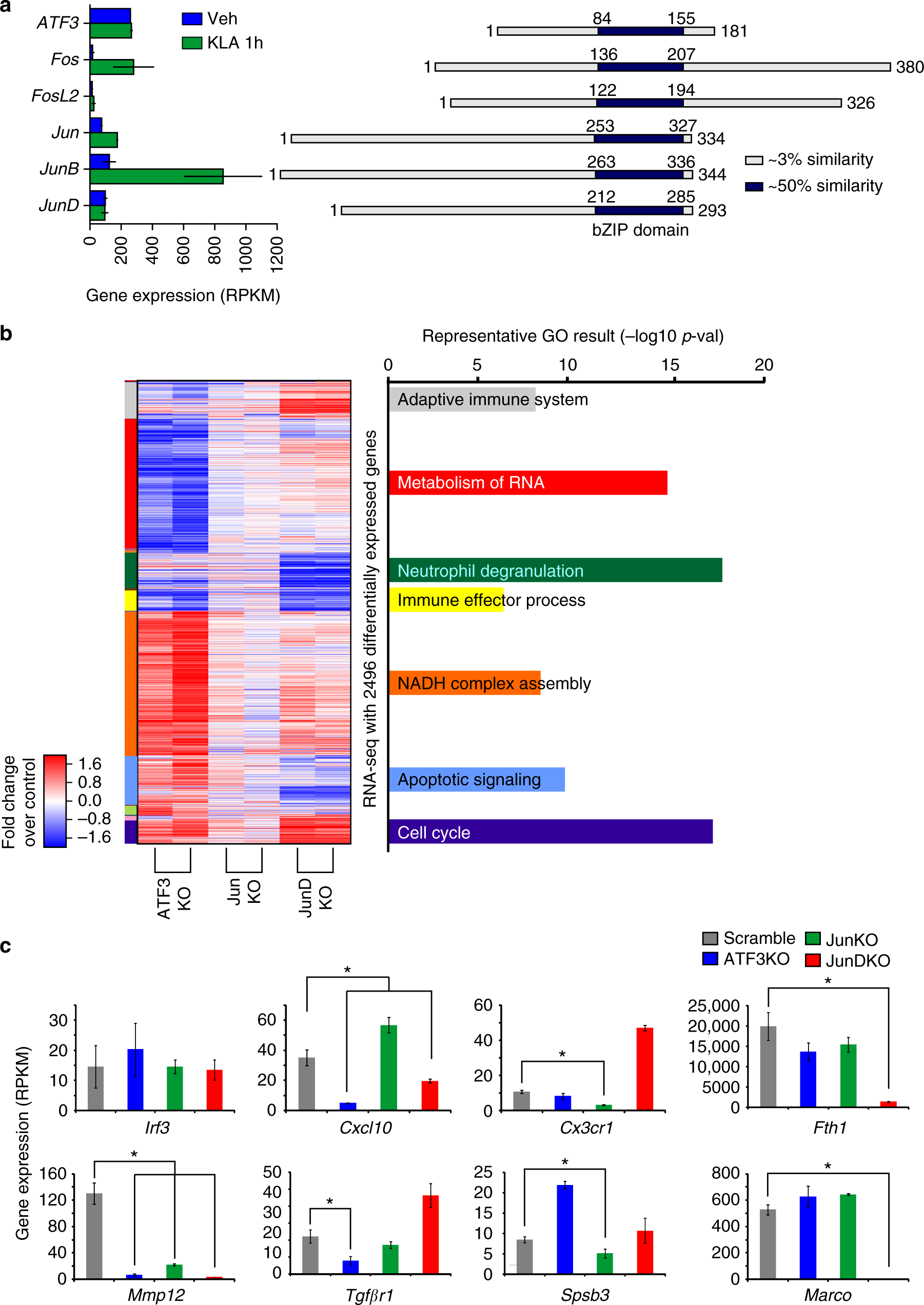
Diverse motif ensembles specify non-redundant DNA binding activities of AP-1 family members in macrophages | Nature Communications
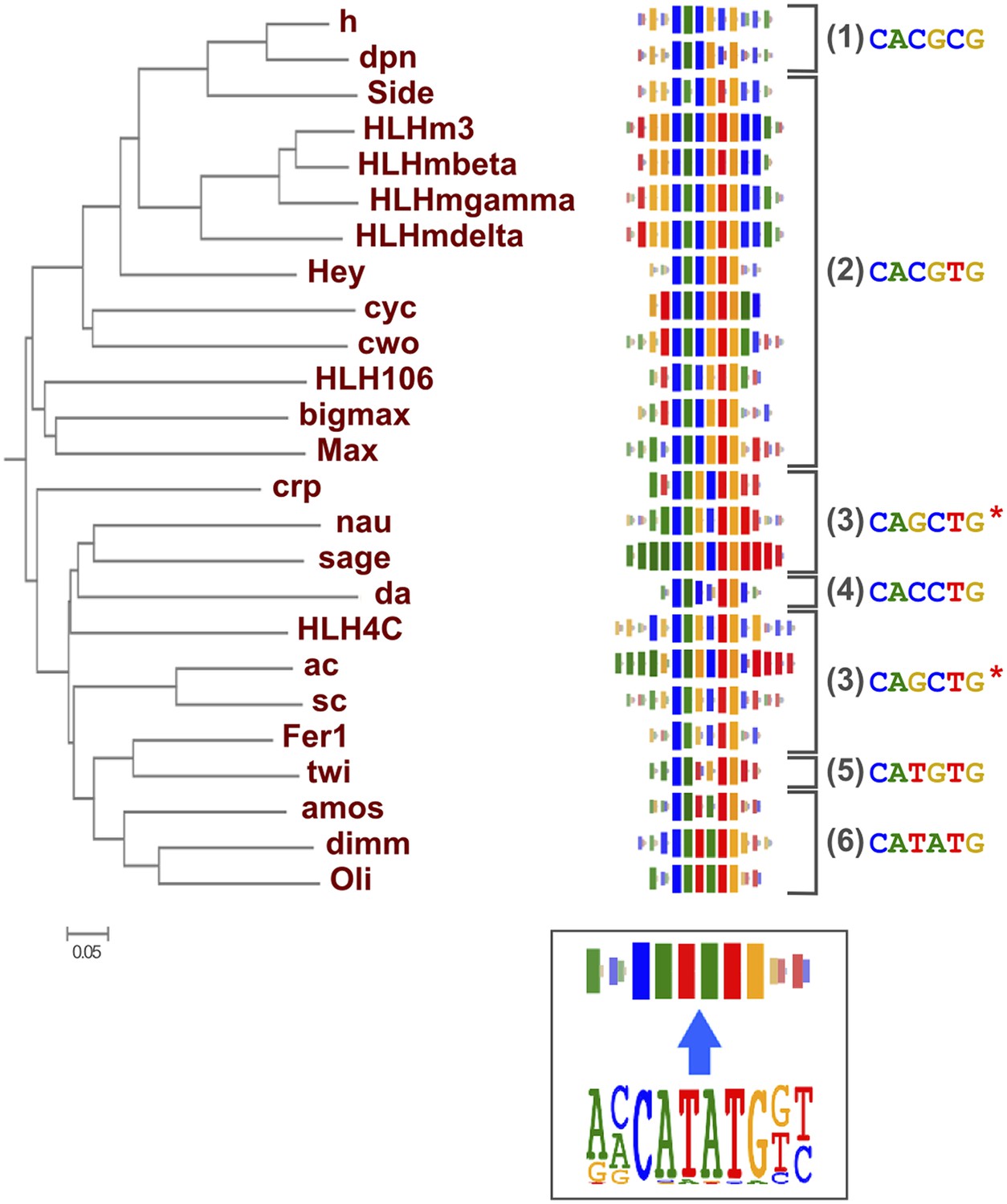
Conservation of transcription factor binding specificities across 600 million years of bilateria evolution | eLife
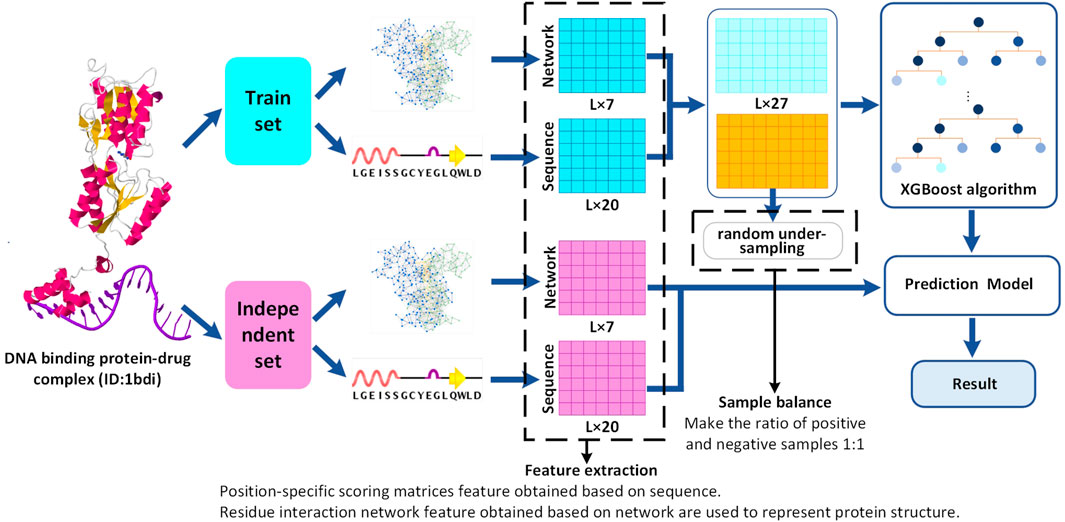
Frontiers | Prediction of DNA-Binding Protein–Drug-Binding Sites Using Residue Interaction Networks and Sequence Feature
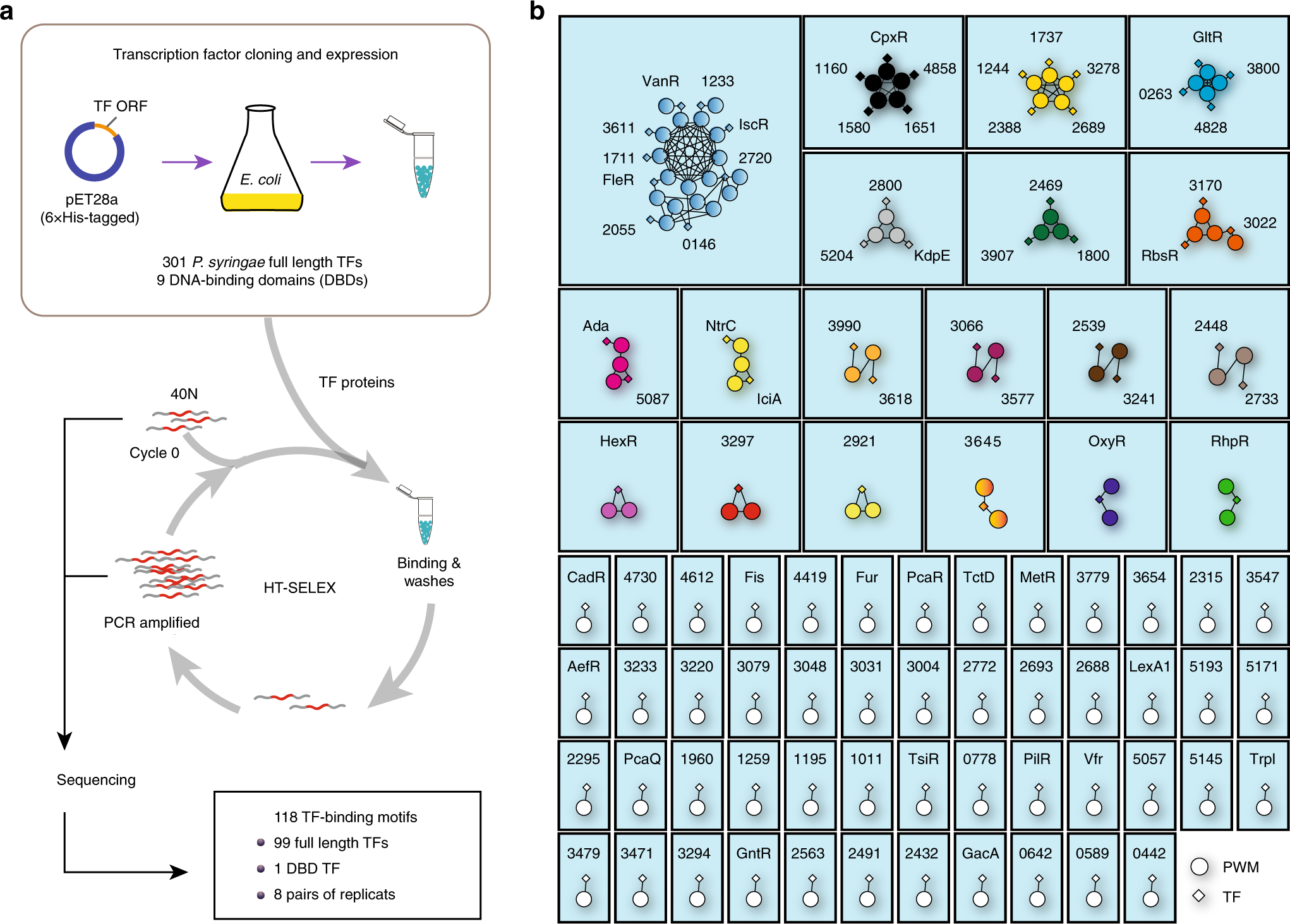
A compendium of DNA-binding specificities of transcription factors in Pseudomonas syringae | Nature Communications
![PDF] DNA Shape Features Improve Transcription Factor Binding Site Predictions In Vivo. | Semantic Scholar PDF] DNA Shape Features Improve Transcription Factor Binding Site Predictions In Vivo. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/5d68bdf0391af25147a48171a54fea49d64d16d4/2-Figure1-1.png)
PDF] DNA Shape Features Improve Transcription Factor Binding Site Predictions In Vivo. | Semantic Scholar