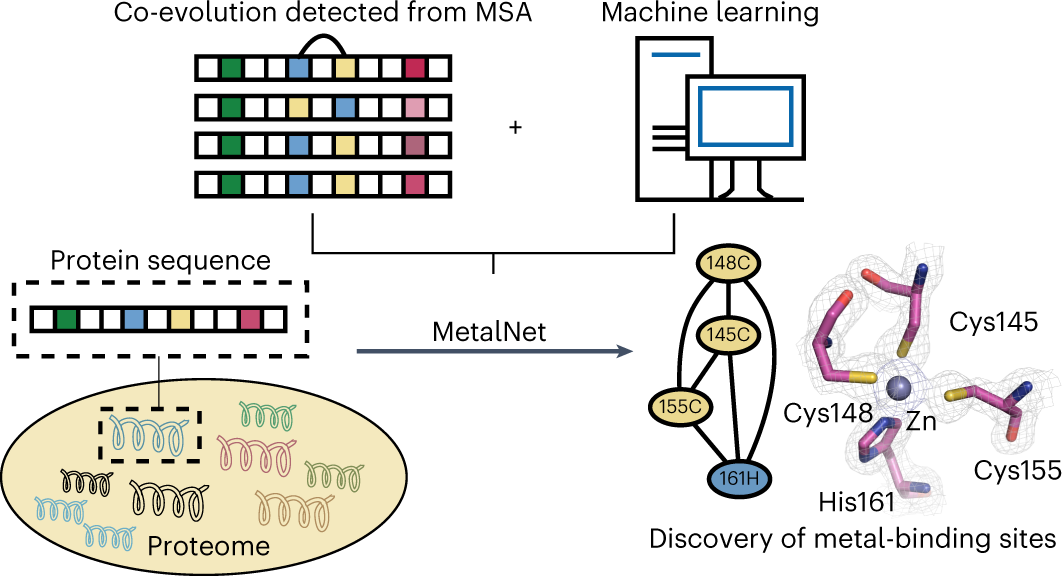
Co-evolution-based prediction of metal-binding sites in proteomes by machine learning | Nature Chemical Biology

Figure 2 from Protein-ligand binding site recognition using complementary binding-specific substructure comparison and sequence profile alignment | Semantic Scholar

Workflow of binding sites detection. Each protein chain submitted would... | Download Scientific Diagram

Can We Rely on Computational Predictions To Correctly Identify Ligand Binding Sites on Novel Protein Drug Targets? Assessment of Binding Site Prediction Methods and a Protocol for Validation of Predicted Binding Sites

Ligand Binding Site Prediction: For the ligand binding sites prediction... | Download Scientific Diagram

Illustration of protein binding site prediction for Targets T0570 (a),... | Download Scientific Diagram
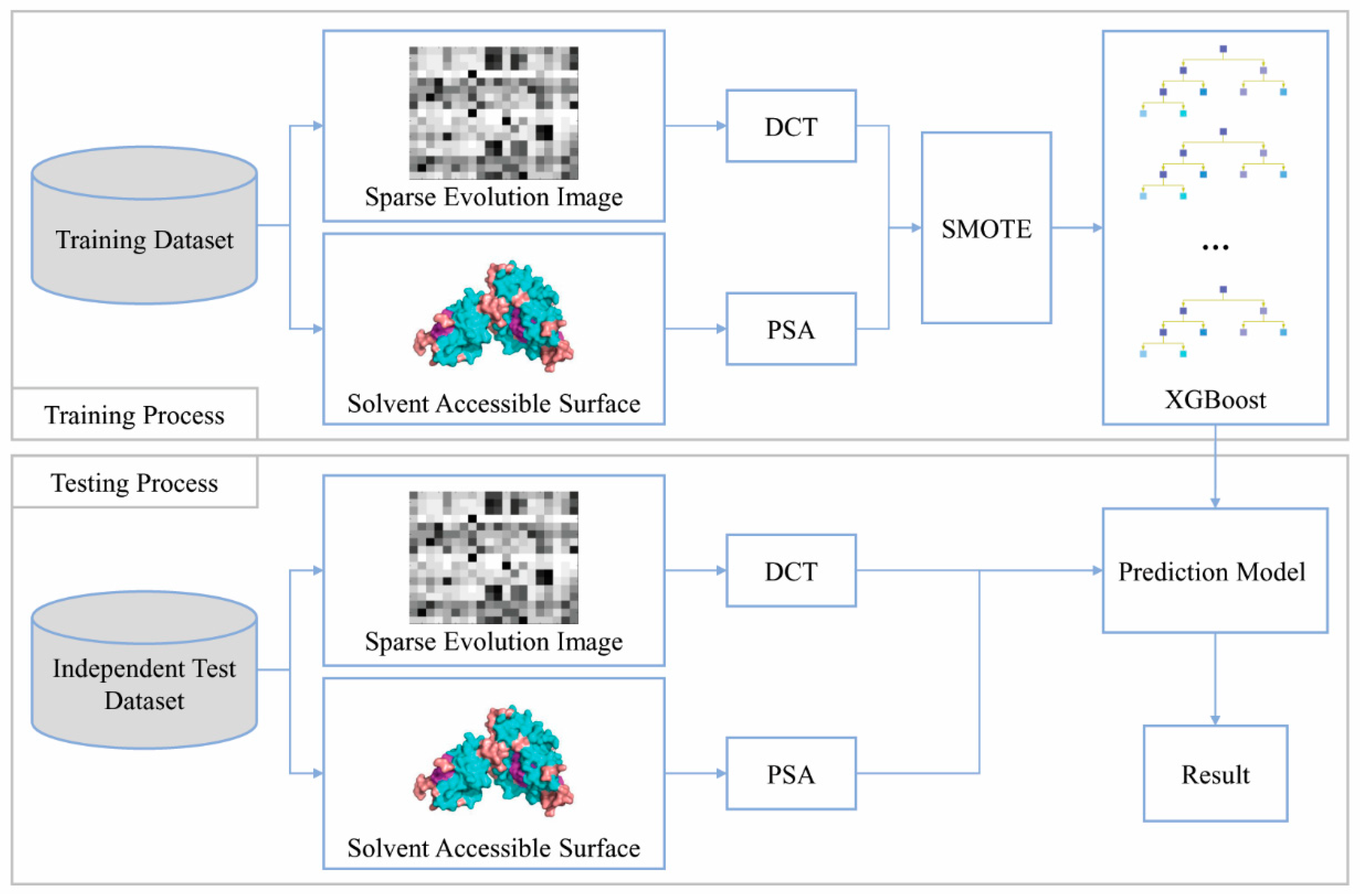
Genes | Free Full-Text | SXGBsite: Prediction of Protein–Ligand Binding Sites Using Sequence Information and Extreme Gradient Boosting

Spatiotemporal identification of druggable binding sites using deep learning | Communications Biology
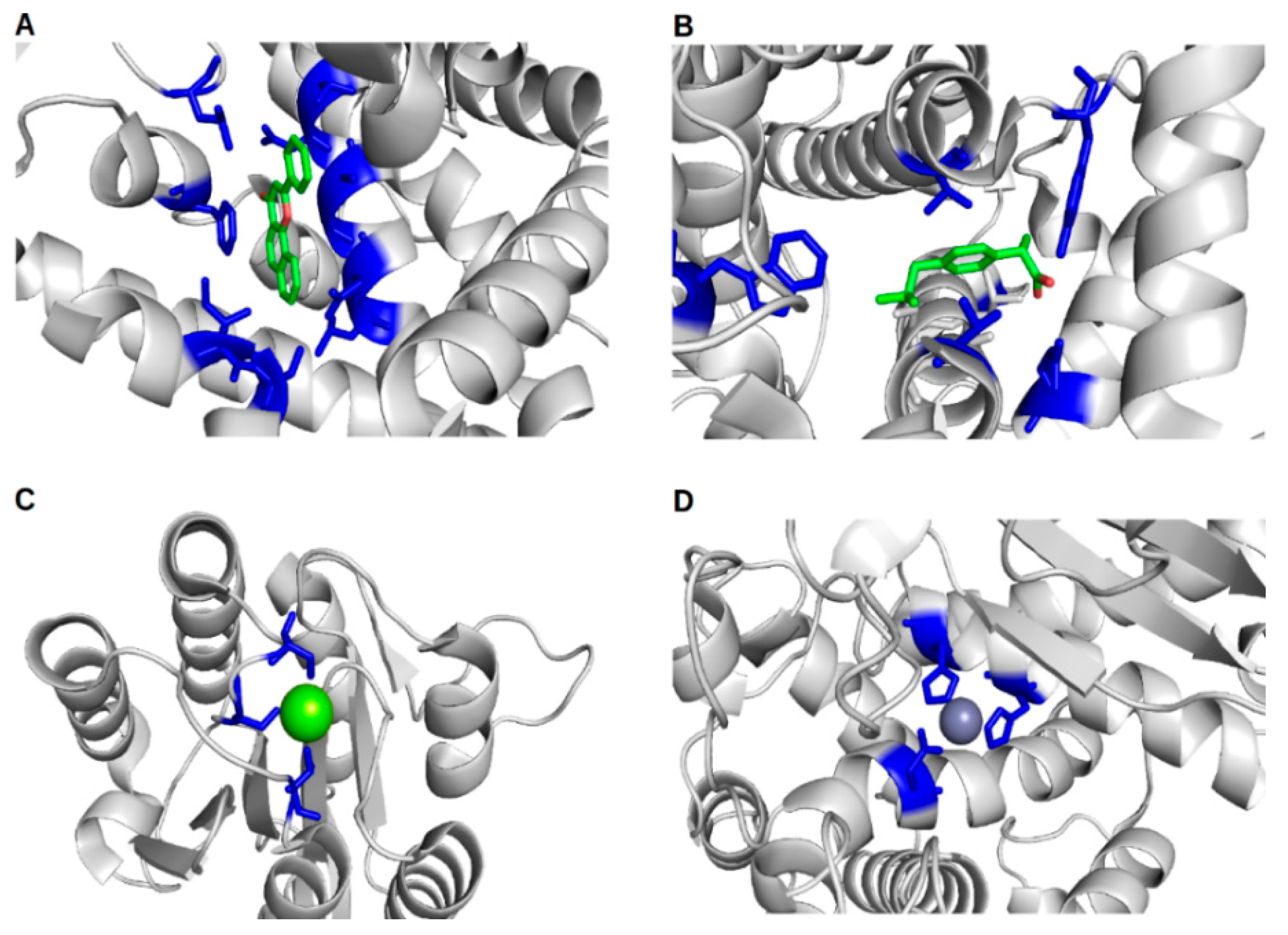
IJMS | Free Full-Text | Proteins and Their Interacting Partners: An Introduction to Protein–Ligand Binding Site Prediction Methods

Recognizing Protein-Ligand Binding Sites by Global Structural Alignment and Local Geometry Refinement - ScienceDirect

Virtual Screening | Target Binding Site Prediction / Identification || Drug Discovery || P2c-3 - YouTube

AI-based prediction of new binding site and virtual screening for the discovery of novel P2X3 receptor antagonists - ScienceDirect

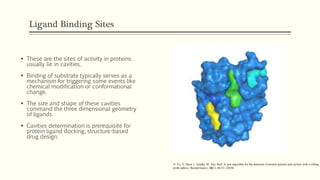

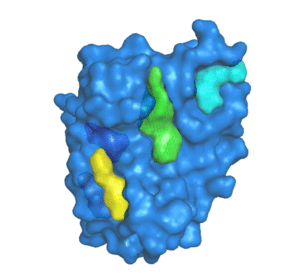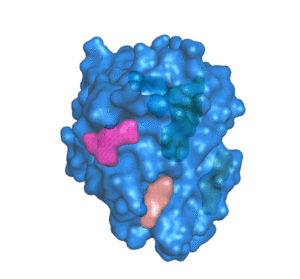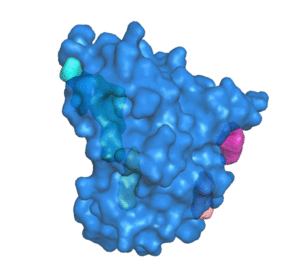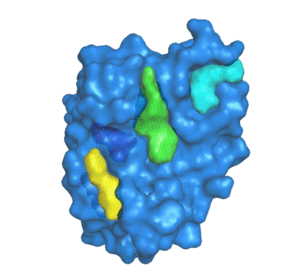


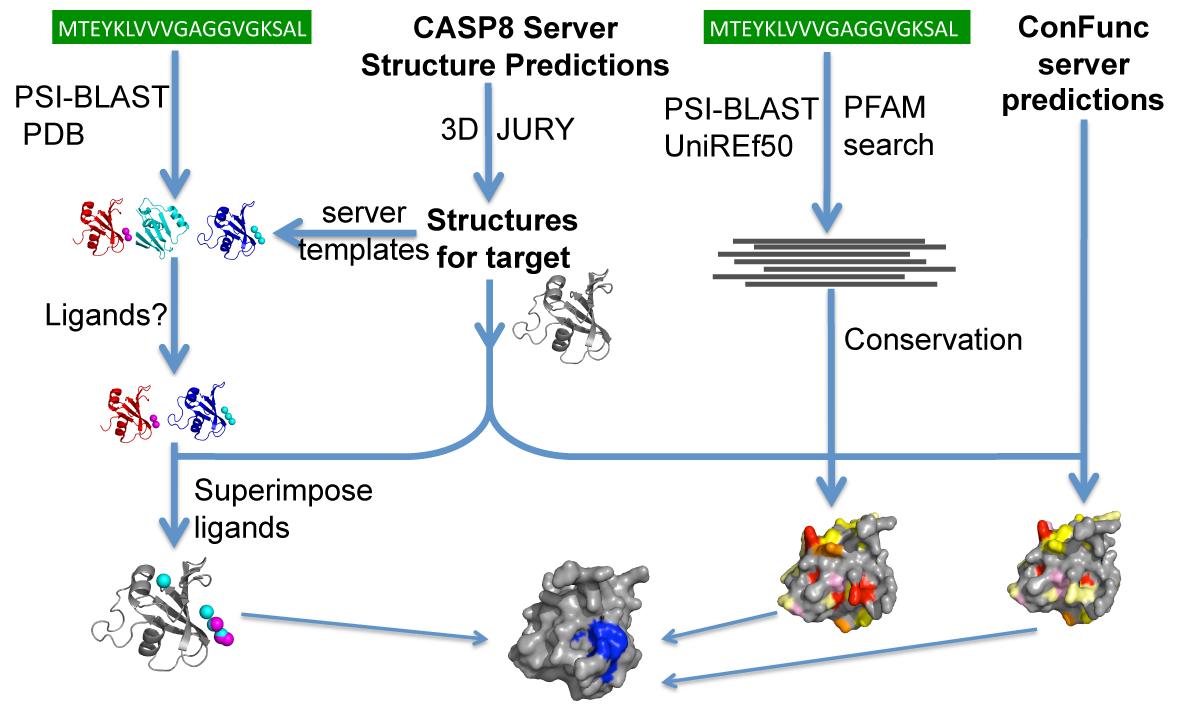
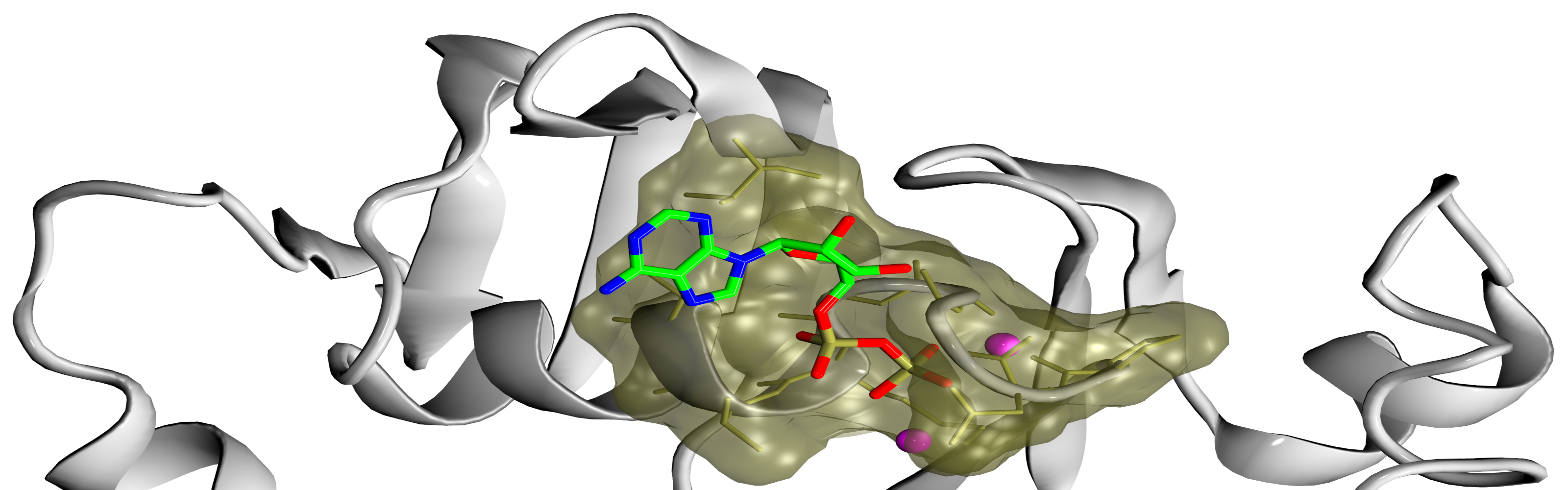
![PDF] The FunFOLD 2 server for the prediction of protein ligand interactions | Semantic Scholar PDF] The FunFOLD 2 server for the prediction of protein ligand interactions | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/c50225f129c18e56d2cbe4352383ce46df326d70/5-Figure1-1.png)
