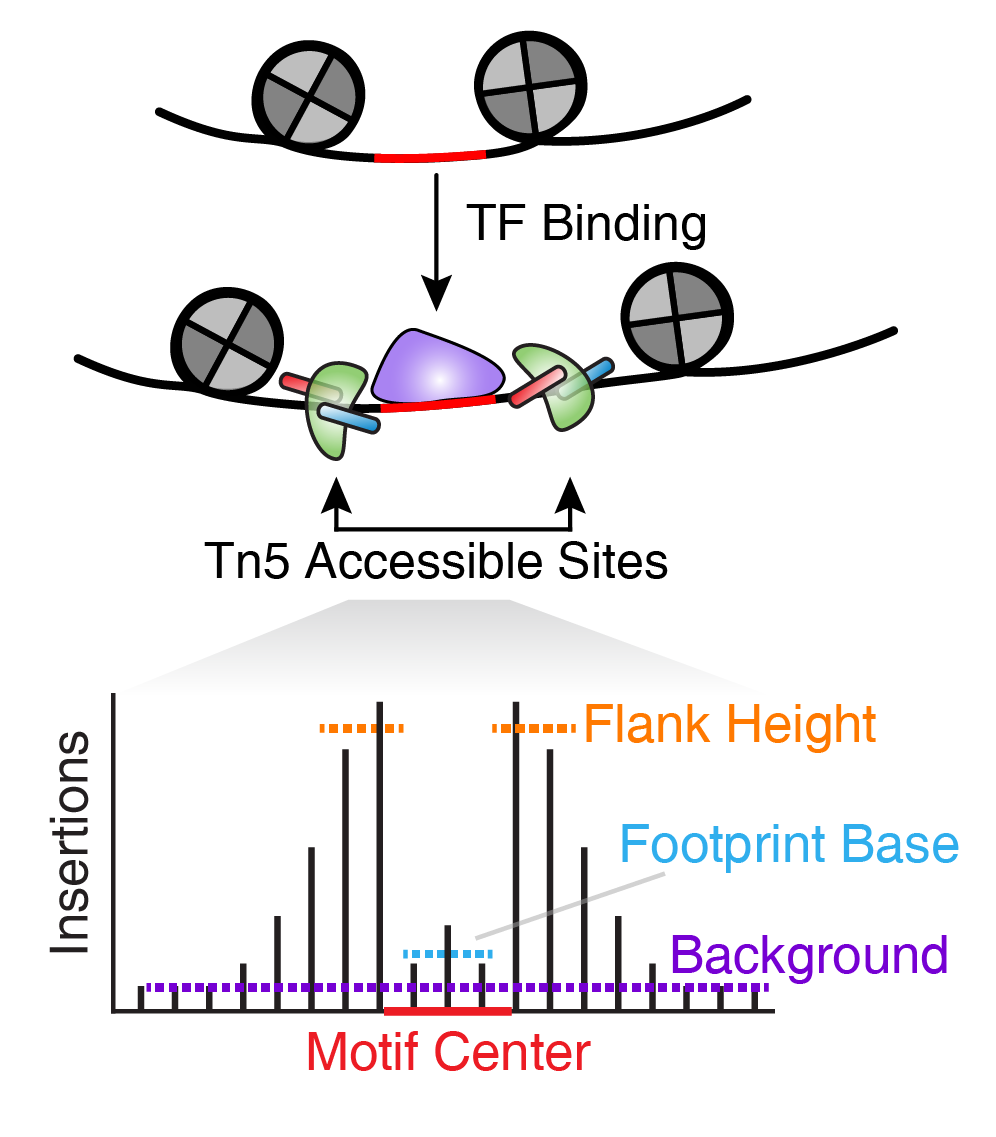
Frontiers | Integrating Peak Colocalization and Motif Enrichment Analysis for the Discovery of Genome-Wide Regulatory Modules and Transcription Factor Recruitment Rules
![PDF] TFBSTools: an R/bioconductor package for transcription factor binding site analysis | Semantic Scholar PDF] TFBSTools: an R/bioconductor package for transcription factor binding site analysis | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/7952b7c68b33972d5f8483cf7742efaed6396357/2-Figure1-1.png)
PDF] TFBSTools: an R/bioconductor package for transcription factor binding site analysis | Semantic Scholar

Prediction of transcription factor binding sites and mutation analysis... | Download Scientific Diagram

Fragment binding site analysis results from FTMAP and identification of... | Download Scientific Diagram
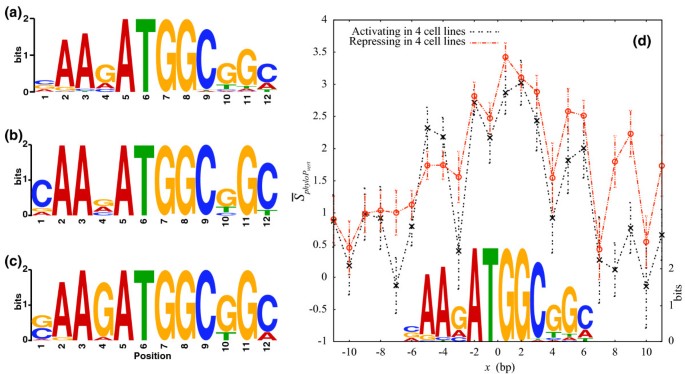
Functional analysis of transcription factor binding sites in human promoters | Genome Biology | Full Text

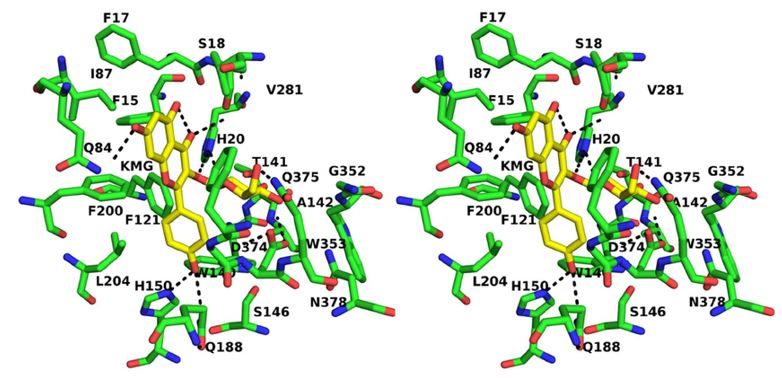
Mapping the protein binding site of the (pro)renin receptor using in silico 3D structural analysis | Hypertension Research

Molecules | Free Full-Text | Identification of Potential Modulators of a Pathogenic G Protein-Gated Inwardly Rectifying K+ Channel 4 Mutant: In Silico Investigation in the Context of Drug Discovery for Hypertension

Identification of transcription factor binding sites on promoter of RNA dependent RNA polymerases (RDRs) and interacting partners of RDR proteins through in silico analysis | Physiology and Molecular Biology of Plants
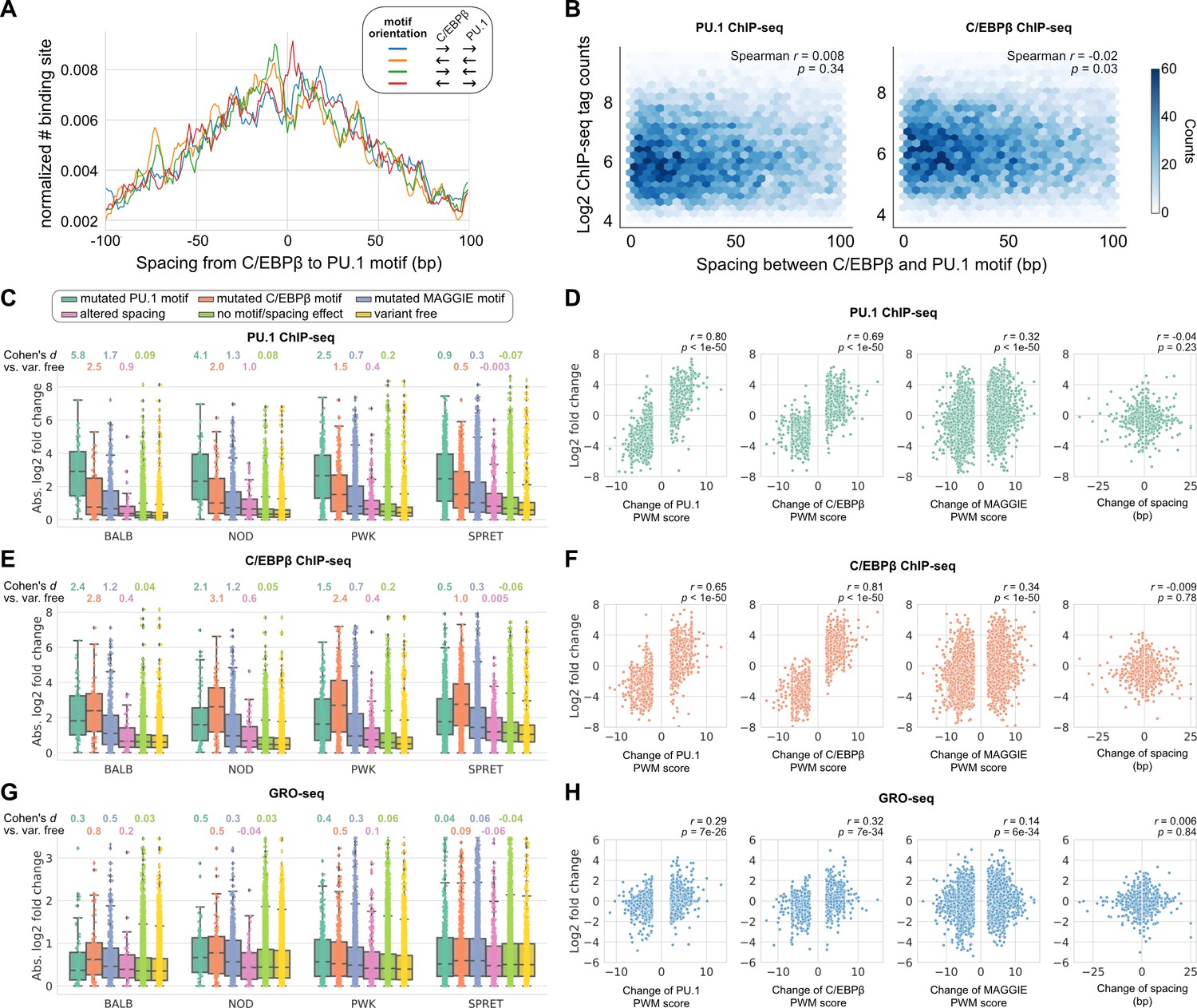
Systematic analysis of naturally occurring insertions and deletions that alter transcription factor spacing identifies tolerant and sensitive transcription factor pairs | eLife