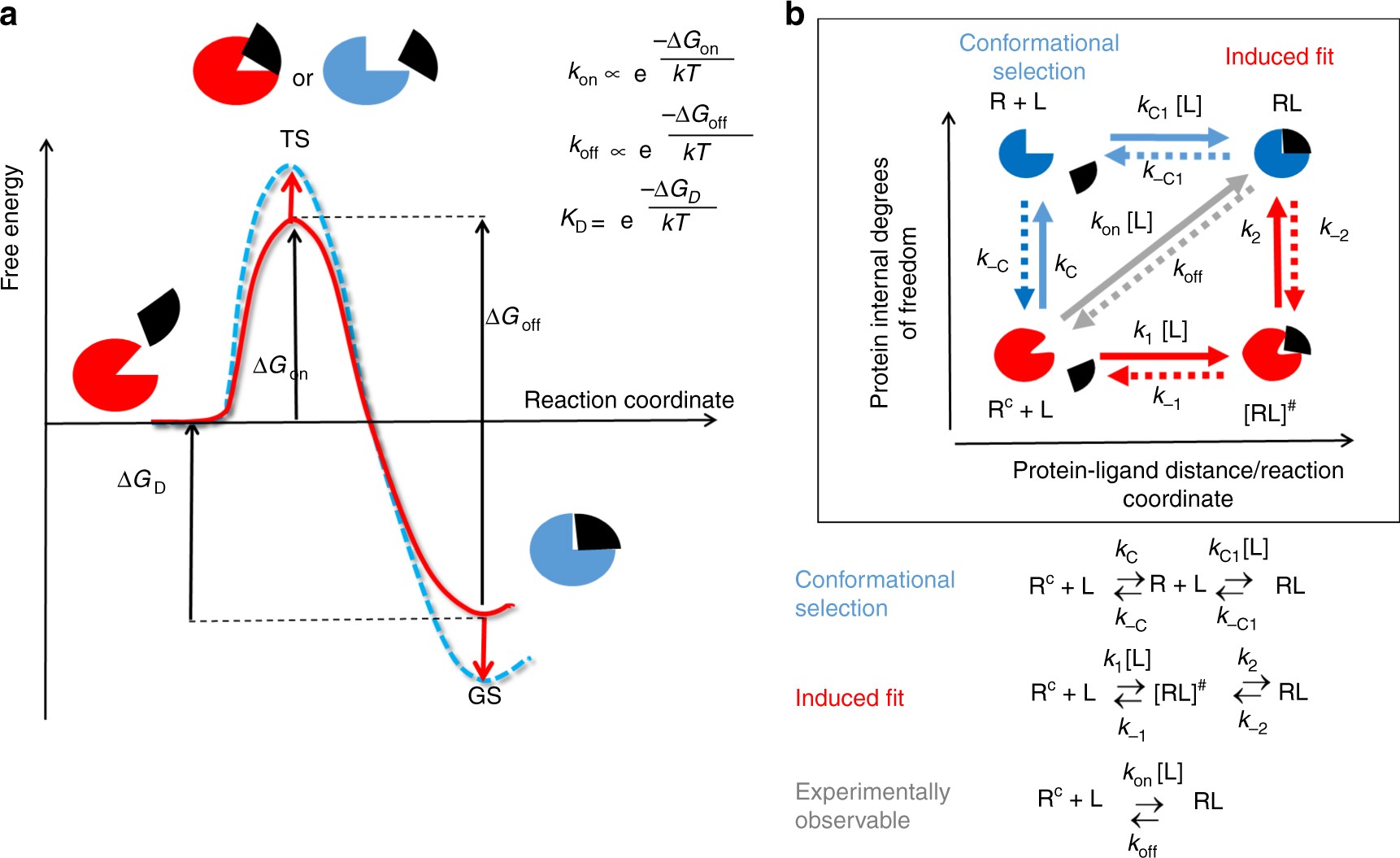
Protein conformational flexibility modulates kinetics and thermodynamics of drug binding | Nature Communications

Relative Binding Enthalpies from Molecular Dynamics Simulations Using a Direct Method | Journal of Chemical Theory and Computation

Binding energy (ΔE), enthalpy (ΔH) and Gibbs free energy (ΔG) of Na 2... | Download Scientific Diagram

Entropy–enthalpy transduction caused by conformational shifts can obscure the forces driving protein–ligand binding | PNAS

Ligand binding free energy and kinetics calculation in 2020 - Limongelli - 2020 - WIREs Computational Molecular Science - Wiley Online Library

Local potential energy: A novel QTAIM tool to quantify the binding energy of classical hydrogen bonds - ScienceDirect

CO Binding Energy is an Incomplete Descriptor of Cu‐Based Catalysts for the Electrochemical CO2 Reduction Reaction - Gao - 2023 - Angewandte Chemie International Edition - Wiley Online Library