
Discovery of a new ATP-binding motif involved in peptidic azoline biosynthesis | Nature Chemical Biology

E2EATP: Fast and High-Accuracy Protein–ATP Binding Residue Prediction via Protein Language Model Embedding,Journal of Chemical Information and Modeling - X-MOL

Full article: A novel sequence-based prediction method for ATP-binding sites using fusion of SMOTE algorithm and random forests classifier

The linker domain of the initiator DnaA contributes to its ATP binding and membrane association in E. coli chromosomal replication | Science Advances

PDF) ATP-binding site as a further application of neural networks to residue level prediction | zulfiqar ahmad - Academia.edu

Full article: Computational prediction and in vitro analysis of the potential ligand binding site within the extracellular ATP receptor, P2K2
Computational Analysis of the Ligand Binding Site of the Extracellular ATP Receptor, DORN1 | PLOS ONE

Comparison of ATP-binding pockets and discovery of homologous recombination inhibitors - ScienceDirect

IJMS | Free Full-Text | Biochemical and Computational Studies of the Interaction between a Glucosamine Derivative, NAPA, and the IKKα Kinase

Exploring the computational methods for protein-ligand binding site prediction - Computational and Structural Biotechnology Journal

Biomolecules | Free Full-Text | Motif2Mol: Prediction of New Active Compounds Based on Sequence Motifs of Ligand Binding Sites in Proteins Using a Biochemical Language Model

Prediction of ATP-binding sites in membrane proteins using a two-dimensional convolutional neural network - ScienceDirect

Prediction of ATP-binding sites in membrane proteins using a two-dimensional convolutional neural network - ScienceDirect

A) 3D position of the sub-pocket in the ATP binding site of PI3K-α and... | Download Scientific Diagram

Conformation transitions of the polypeptide-binding pocket support an active substrate release from Hsp70s | Nature Communications

Active Site Sequence Representations of Human Kinases Outperform Full Sequence Representations for Affinity Prediction and Inhibitor Generation: 3D Effects in a 1D Model | Journal of Chemical Information and Modeling

ATPbind: Accurate Protein–ATP Binding Site Prediction by Combining Sequence-Profiling and Structure-Based Comparisons | Journal of Chemical Information and Modeling
ATPsite: sequence-based prediction of ATP-binding residues. : Chen, Ke : Free Download, Borrow, and Streaming : Internet Archive

Identification and Structural Characterization of the ATP/ADP-Binding Site in the Hsp90 Molecular Chaperone: Cell
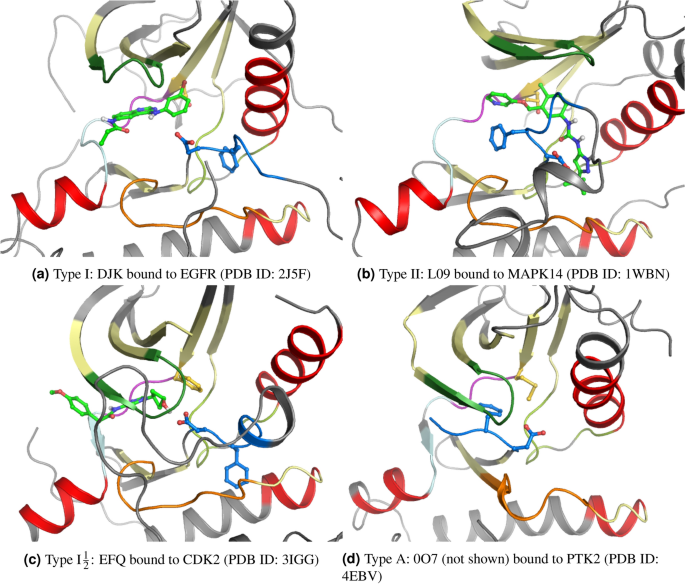
Prediction of kinase inhibitors binding modes with machine learning and reduced descriptor sets | Scientific Reports

TargetATPsite: A template‐free method for ATP‐binding sites prediction with residue evolution image sparse representation and classifier ensemble - Yu - 2013 - Journal of Computational Chemistry - Wiley Online Library


